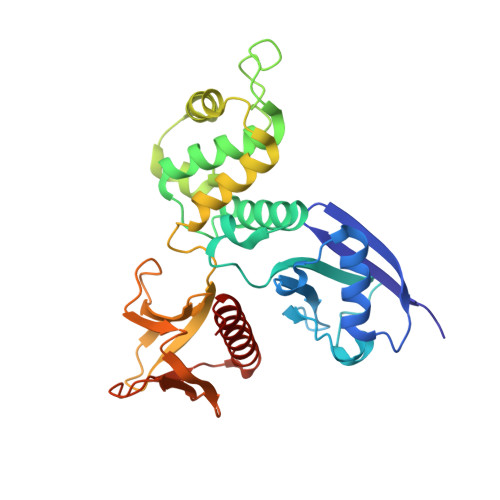
Structure of the Active N-terminal Domain of Ezrin. CONFORMATIONAL AND MOBILITY CHANGES IDENTIFY KEYSTONE INTERACTIONS.
Smith, W.J., Nassar, N., Bretscher, A.P., Cerione, R.A., Karplus, P.A.(2003) J Biological Chem 278: 4949-4956
- PubMed: 12429733 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M210601200
- Primary Citation Related Structures:
1NI2 - PubMed Abstract:
Ezrin is a member of the ERM (ezrin, radixin, moesin) family of proteins that cross-link the actin cytoskeleton to the plasma membrane and also may function in signaling cascades that regulate the assembly of actin stress fibers. Here, we report a crystal structure for the free (activated) FERM domain (residues 2-297) of recombinant human ezrin at 2.3 A resolution. Structural comparison among the dormant moesin FERM domain structure and the three known active FERM domain structures (radixin, moesin, and now ezrin) allows the clear definition of regions that undergo structural changes during activation. The key regions affected are residues 135-150 and 155-180 in lobe F2 and residues 210-214 and 235-267 in lobe F3. Furthermore, we show that a large increase in the mobilities of lobes F2 and F3 accompanies activation, suggesting that their integrity is compromised. This leads us to propose a new concept that we refer to as keystone interactions. Keystone interactions occur when one protein (or protein part) contributes residues that allow another protein to complete folding, meaning that it becomes an integral part of the structure and would rarely dissociate. Such interactions are well suited for long-lived cytoskeletal protein interactions. The keystone interactions concept leads us to predict two specific docking sites within lobes F2 and F3 that are likely to bind target proteins.
- Department of Chemistry, Cornell University, Ithaca, New York 14853, USA.
Organizational Affiliation: