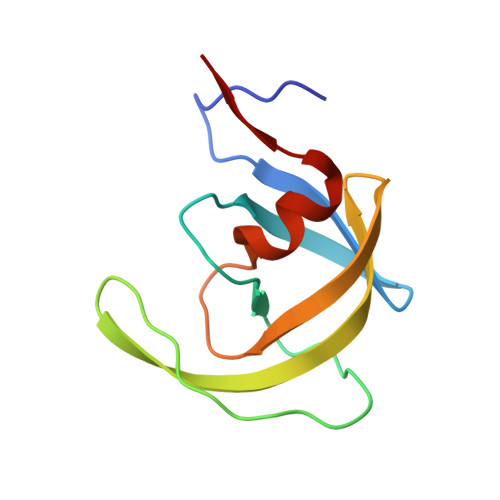
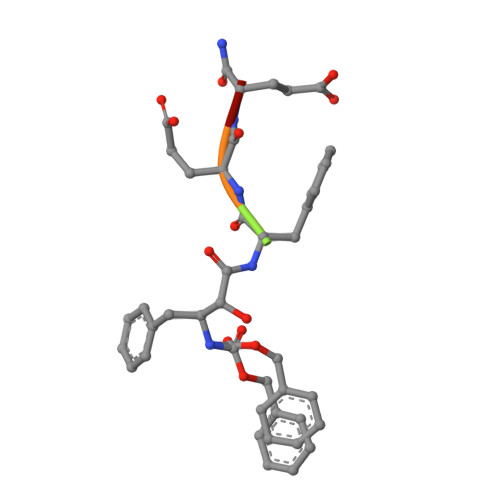
A Phenylnorstatine Inhibitor Binding to HIV-1 Protease: Geometry, Protonation, and Subsite-Pocket Interactions Analyzed at Atomic Resolution
Brynda, J., Rezacova, P., Fabry, M., Horejsi, M., Stouracova, R., Sedlacek, J., Soucek, M., Hradilek, M., Lepsik, M., Konvalinka, J.(2004) J Med Chem 47: 2030-2036
- PubMed: 15056001 Search on PubMed
- DOI: https://doi.org/10.1021/jm031105q
- Primary Citation Related Structures:
1NH0 - PubMed Abstract:
The X-ray structure of a complex of HIV-1 protease (PR) with a phenylnorstatine inhibitor Z-Pns-Phe-Glu-Glu-NH(2) has been determined at 1.03 A, the highest resolution so far reported for any HIV PR complex. The inhibitor shows subnanomolar K(i) values for both the wild-type PR and the variant representing one of the most common mutations linked to resistance development. The structure comprising the phenylnorstatine moiety of (2R,3S)-chirality displays a unique pattern of hydrogen bonding to the two catalytic aspartate residues. This high resolution makes it possible to assess the donor and acceptor relations of this hydrogen bonding and to indicate a proton shared by the two catalytic residues. A structural mechanism for the unimpaired inhibition of the protease Val82Ala mutant is also suggested, based on energy calculations and analyses.
- Institute of Molecular Genetics, Academy of Sciences of the Czech Republic, Flemingovo nám. 2, 16637 Prague 6, Czech Republic. brynda@img.cas.cz
Organizational Affiliation: