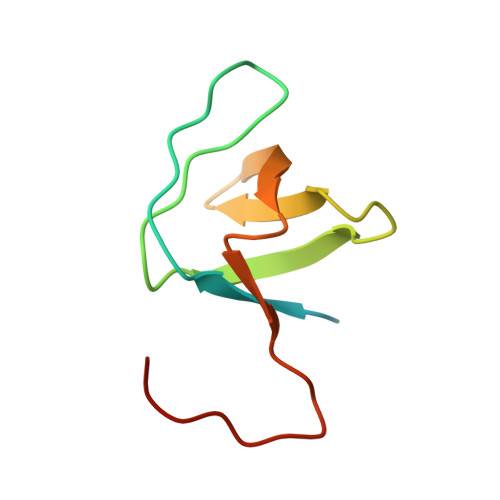
Rapid purification and crystal structure analysis of a small protein carrying two terminal affinity tags
Mueller, U., Buessow, K., Diehl, A., Bartl, F.J., Niesen, F.H., Nyarsik, L., Heinemann, U.(2003) J Struct Funct Genomics 4: 217-225
- PubMed: 15185962 Search on PubMed
- DOI: https://doi.org/10.1023/b:jsfg.0000016119.50040.a3
- Primary Citation Related Structures:
1NEG - PubMed Abstract:
Small peptide tags are often fused to proteins to allow their affinity purification in high-throughput structure analysis schemes. To assess the compatibility of small peptide tags with protein crystallization and to examine if the tags alter the three-dimensional structure, the N-terminus of the chicken alpha-spectrin SH3 domain was labeled with a His6 tag and the C-terminus with a StrepII tag. The resulting protein, His6-SH3-StrepII, consists of 83 amino-acid residues, 23 of which originate from the tags. His6-SH3-StrepII is readily purified by dual affinity chromatography, has very similar biophysical characteristics as the untagged protein domain and crystallizes readily from a number of sparse-matrix screen conditions. The crystal structure analysis at 2.3 A resolution proves native-like structure of His6-SH3-StrepII and shows the entire His6 tag and part of the StrepII tag to be disordered in the crystal. Obviously, the fused affinity tags did not interfere with crystallization and structure analysis and did not change the protein structure. From the extreme case of His6-SH3-StrepII, where affinity tags represent 27% of the total fusion protein mass, we extrapolate that protein constructs with N- and C-terminal peptide tags may lend themselves to biophysical and structural investigations in high-throughput regimes.
- Institut für Chemie/Kristallographie, Freie Universität Berlin, Takustr. 6, D-14195 Berlin, Germany.
Organizational Affiliation: