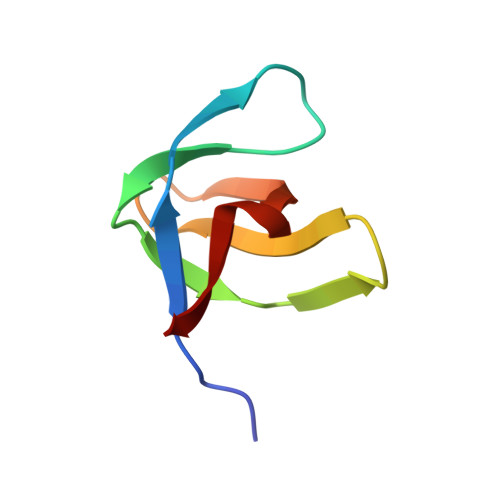
SH3 in muscles: solution structure of the SH3 domain from nebulin.
Politou, A.S., Millevoi, S., Gautel, M., Kolmerer, B., Pastore, A.(1998) J Mol Biology 276: 189-202
- PubMed: 9514727
- DOI: https://doi.org/10.1006/jmbi.1997.1521
- Primary Citation Related Structures:
1ARK, 1NEB - PubMed Abstract:
The huge modular protein nebulin is located in the thin filament of striated muscle in vertebrates and is thought to bind and stabilize F-actin. The C-terminal part of human nebulin is anchored in the sarcomeric Z-disk and contains an SH3 domain, the first of such motifs to be identified in a myofibrillar protein. We have determined the nebulin SH3 sequence from several species and found it strikingly conserved. We have also shown that the SH3 transcripts are constitutively expressed in skeletal muscle tissues. As the first step towards a molecular understanding of nebulin's cellular role we have determined the three-dimensional structure of the human nebulin SH3 domain in solution by nuclear magnetic resonance (NMR) spectroscopy and compared it with other known SH3 structures. The nebulin SH3 domain has a well-defined structure in solution with a typical SH3 topology, consisting of a beta-sandwich of two triple-stranded, antiparallel beta-sheets arranged at right angles to each other and of a single turn of a 310-helix. An additional double-stranded antiparallel beta-sheet in the RT loop bends over the beta-sandwich. The derived structure reveals a remarkable similarity with a distinct subset of SH3 domains, especially in the structural features of the exposed hydrophobic patch that is thought to be the site of interaction with polyproline ligands. On the basis of this similarity, we have modelled the interaction with an appropriate polyproline ligand and attempted to delineate the characteristics of the physiological SH3-binding partner in the Z-disk. Our results represent the first step in reconstructing the structure of nebulin and are expected to contribute to our understanding of nebulin's functional role in myofibrillar assembly.
- Chemistry Department University of Crete, Heraklion, Greece.
Organizational Affiliation: