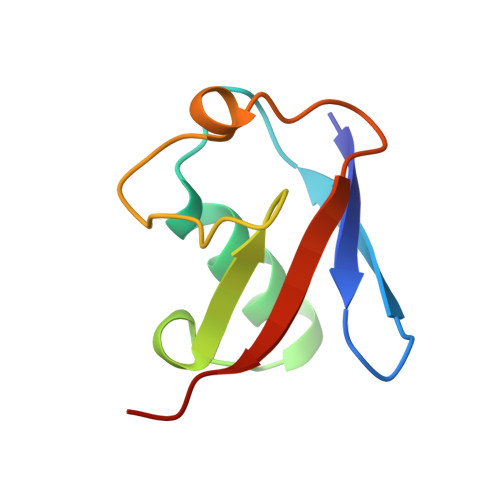
Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes.
Whitby, F.G., Xia, G., Pickart, C.M., Hill, C.P.(1998) J Biological Chem 273: 34983-34991
- PubMed: 9857030 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.273.52.34983
- Primary Citation Related Structures:
1NDD - PubMed Abstract:
The NEDD8/Rub1 class of ubiquitin-like proteins has been implicated in progression of the cell cycle from G1 into S phase. These molecules undergo a metabolism that parallels that of ubiquitin and involves specific interactions with many different proteins. We report here the crystal structure of recombinant human NEDD8 refined at 1.6-A resolution to an R factor of 21.9%. As expected from the high sequence similarity (57% identical), the NEDD8 structure closely resembles that reported previously for ubiquitin. We also show that recombinant human NEDD8 protein is activated, albeit inefficiently, by the ubiquitin-activating (E1) enzyme and that NEDD8 can be transferred from E1 to the ubiquitin conjugating enzyme E2-25K. E2-25K adds NEDD8 to a polyubiquitin chain with an efficiency similar to that of ubiquitin. A chimeric tetramer composed of three ubiquitins and one histidine-tagged NEDD8 binds to the 26 S proteasome with an affinity similar to that of tetraubiquitin. Seven residues that differ from the corresponding residues in ubiquitin, but are conserved between NEDD8 orthologs, are candidates for mediating interactions with NEDD8-specific partners. One such residue, Ala-72 (Arg in ubiquitin), is shown to perform a key role in selecting against reaction with the ubiquitin E1 enzyme, thereby acting to prevent the inappropriate diversion of NEDD8 into ubiquitin-specific pathways.
- Department of Biochemistry, University of Utah, Salt Lake City, Utah 84132, USA.
Organizational Affiliation: