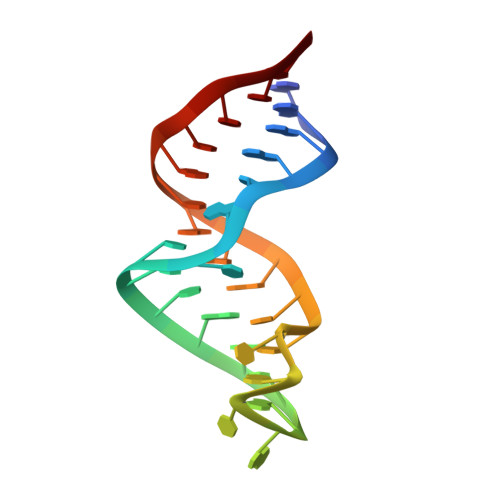
Refined Solution Structure of the Iron-responsive Element RNA Using Residual Dipolar Couplings
McCallum, S.A., Pardi, A.(2003) J Mol Biology 326: 1037-1050
- PubMed: 12589752 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)01431-6
- Primary Citation Related Structures:
1NBR - PubMed Abstract:
The iron-responsive element (IRE) is a 30nt RNA motif located in the non-coding regions of mRNAs of proteins involved in iron regulation. In humans, the IRE plays a direct role in the control of iron levels by post-transcriptional regulation of the ferritin and transferrin receptor proteins through highly specific recognition by IRE-binding proteins. The IRE fold is representative of many RNA motifs that contain helical domains separated by a bulge or internal loop. The global structures of such extended multi-domain RNAs are not well defined by conventional NMR-distance and torsion angle structural restraints. Residual dipolar couplings (RDCs) are employed here to better define the global structure of the IRE RNA in solution. RDCs contain valuable long-range structural information that compliments the short-range structural data derived from standard NOE-distance and torsion angle restraints. Several approaches for estimating alignment tensor parameters and incorporating RDCs into RNA structure determinations are compared. Both the local and global structure of the IRE are improved significantly by refinement with RDCs. These RDC refinements provide insight on the conformational dynamics of the IRE. These studies highlight some issues that need to be addressed when incorporating RDCs in solution structure determinations of nucleic acids. The approach used here should prove valuable for structure determinations of various multi-domain systems.
- Department of Chemistry and Biochemistry, 215 UCB, University of Colorado, Boulder, CO 80309-0215, USA.
Organizational Affiliation: