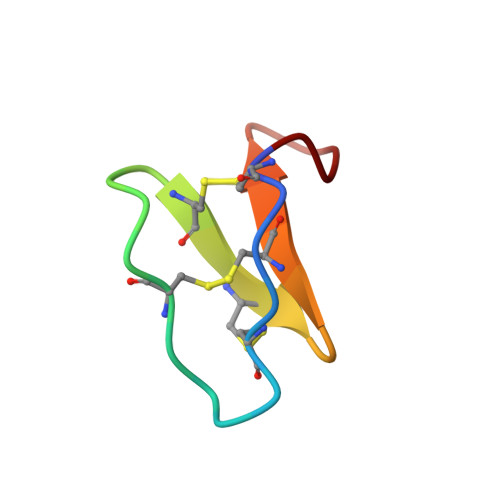
Twists, Knots, and Rings in Proteins. STRUCTURAL DEFINITION OF THE CYCLOTIDE FRAMEWORK
Rosengren, K.J., Daly, N.L., Plan, M.R., Waine, C., Craik, D.J.(2003) J Biological Chem 278: 8606-8616
- PubMed: 12482868 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M211147200
- Primary Citation Related Structures:
1NB1, 1NBJ - PubMed Abstract:
In recent years an increasing number of miniproteins containing an amide-cyclized backbone have been discovered. The cyclotide family is the largest group of such proteins and is characterized by a circular protein backbone and six conserved cysteine residues linked by disulfide bonds in a tight core of the molecule. These form a cystine knot in which an embedded ring formed by two of the disulfide bonds and the connecting backbone segment is threaded by a third disulfide bond. In the current study we have undertaken high resolution structural analysis of two prototypic cyclotides, kalata B1 and cycloviolacin O1, to define the role of the conserved residues in the sequence. We provide the first comprehensive analysis of the topological features in this unique family of proteins, namely rings (a circular backbone), twists (a cis-peptide bond in the Möbius cyclotides) and knots (a knotted arrangement of the disulfide bonds).
- Institute for Molecular Bioscience, Australian Research Council Special Research Centre for Functional and Applied Genomics, University of Queensland, Brisbane, Australia.
Organizational Affiliation: