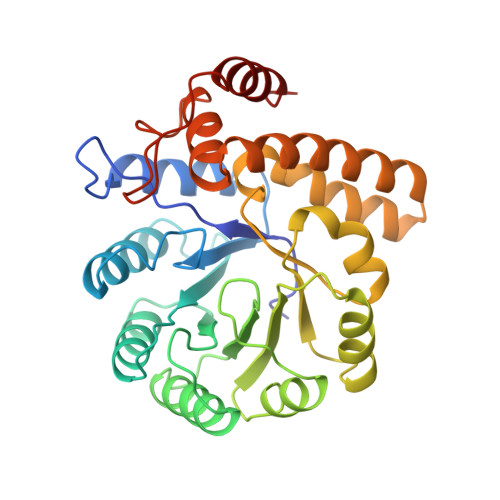
The three-dimensional structure of N-acetylneuraminate lyase from Escherichia coli.
Izard, T., Lawrence, M.C., Malby, R.L., Lilley, G.G., Colman, P.M.(1994) Structure 2: 361-369
- PubMed: 8081752 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00038-1
- Primary Citation Related Structures:
1NAL - PubMed Abstract:
N-acetylneuraminate lyase catalyzes the cleavage of N-acetylneuraminic acid (sialic acid) to form pyruvate and N-acetyl-D-mannosamine. The enzyme plays an important role in the regulation of sialic acid metabolism in bacteria. The reverse reaction can be exploited for the synthesis of sialic acid and some of its derivatives. The structure of the enzyme from Escherichia coli has been determined to 2.2 A resolution by X-ray crystallography. The enzyme is shown to be a tetramer, in which each subunit consists of an alpha/beta-barrel domain followed by a carboxy-terminal extension of three alpha-helices. The active site of the enzyme is tentatively identified as a pocket at the carboxy-terminal end of the eight-stranded beta-barrel. Lys165 lies within this pocket and is probably the reactive residue which forms a Schiff base intermediate with the substrate. The sequence of N-acetylneuraminate lyase has similarities to those of dihydrodipicolinate synthase and MosA (an enzyme implicated in rhizopine synthesis) suggesting that these last two enzymes share a similar structure to N-acetylneuraminate lyase.
- Biomolecular Research Institute, Parkville, Australia.
Organizational Affiliation: