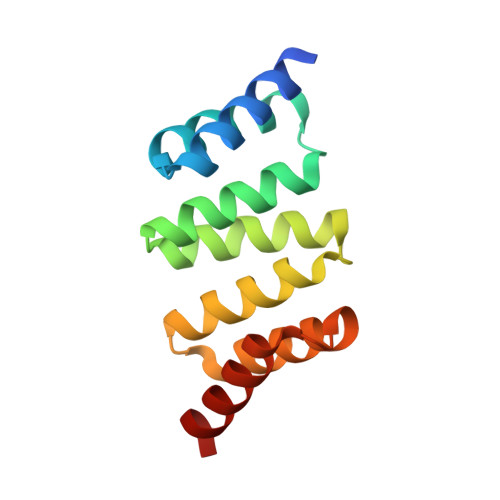
Design of Stable alpha-Helical Arrays from an Idealized TPR Motif
Main, E., Xiong, Y., Cocco, M., D'Andrea, L., Regan, L.(2003) Structure 11: 497-508
- PubMed: 12737816 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00076-5
- Primary Citation Related Structures:
1NA0, 1NA3 - PubMed Abstract:
The tetratricopeptide repeat (TPR) is a 34-amino acid alpha-helical motif that occurs in over 300 different proteins. In the different proteins, three to sixteen or more TPR motifs occur in tandem arrays and function to mediate protein-protein interactions. The binding specificity of each TPR protein is different, although the underlying structural motif is the same. Here we describe a statistical approach to the design of an idealized TPR motif. We present the high-resolution X-ray crystal structures (to 1.55 and 1.6 A) of designed TPR proteins and describe their solution properties and stability. A detailed analysis of these structures provides an understanding of the TPR motif, how it is repeated to give helical arrays with different superhelical twists, and how a very stable framework may be constructed for future functional designs.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520, USA.
Organizational Affiliation: