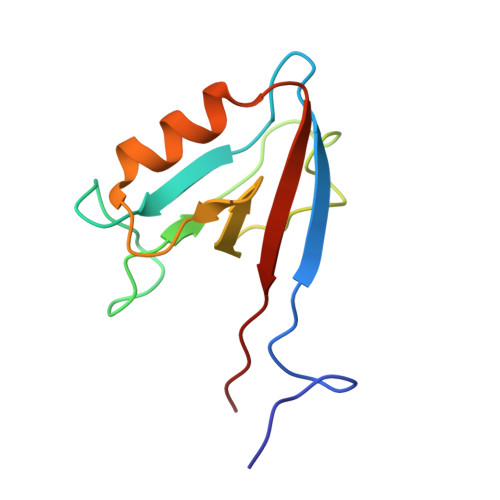
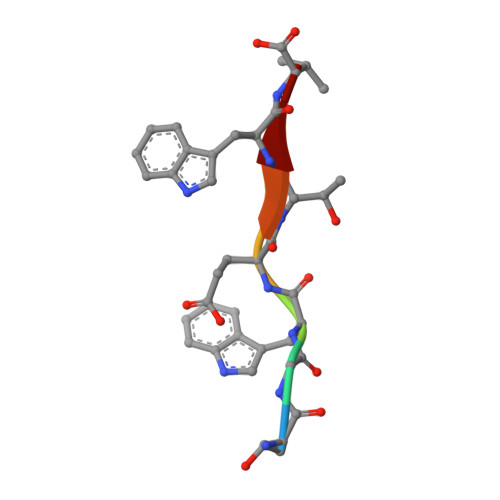
Origins of PDZ domain ligand specificity. Structure determination and mutagenesis of the Erbin PDZ domain.
Skelton, N.J., Koehler, M.F.T., Zobel, K., Wong, W.L., Yeh, S., Pisabarro, M.T., Yin, J.P., Lasky, L.A., Sidhu, S.S.(2003) J Biological Chem 278: 7645-7654
- PubMed: 12446668 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M209751200
- Primary Citation Related Structures:
1N7T - PubMed Abstract:
The LAP (leucine-rich repeat and PDZ-containing) family of proteins play a role in maintaining epithelial and neuronal cell size, and mutation of these proteins can have oncogenic consequences. The LAP protein Erbin has been implicated previously in a number of cellular activities by virtue of its PDZ domain-dependent association with the C termini of both ERB-B2 and the p120-catenins. The present work describes the NMR structure of Erbin PDZ in complex with a high affinity peptide ligand and includes a comprehensive energetic analysis of both the ligand and PDZ domain side chains responsible for binding. C-terminal phage display has been used to identify preferred ligands, whereas binding affinity measurements provide precise details of the energetic importance of each ligand side chain to binding. Alanine and homolog scanning mutagenesis (in a combinatorial phage display format) identifies Erbin side chains that make energetically important contacts with the ligand. The structure of a phage-optimized peptide (Ac-TGW(-4)ETW(-1)V; IC(50) = approximately 0.15 microm) in complex with Erbin PDZ provides a structural context to understand the binding energetics. In particular, the very favorable interactions with Trp(-1) are not Erbin side chain-mediated (and therefore may be generally applicable to many PDZ domains), whereas the beta2-beta3 loop provides a binding site for the Trp(-4) side chain (specific to Erbin because it has an unusually long loop). These results contribute to a growing appreciation for the importance of at least five ligand C-terminal side chains in determining PDZ domain binding energy and highlight the mechanisms of ligand discrimination among the several hundred PDZ domains present in the human genome.
- Department of Protein Engineering, Genentech, Inc., South San Francisco, California 94080, USA. skelly@gene.com
Organizational Affiliation: