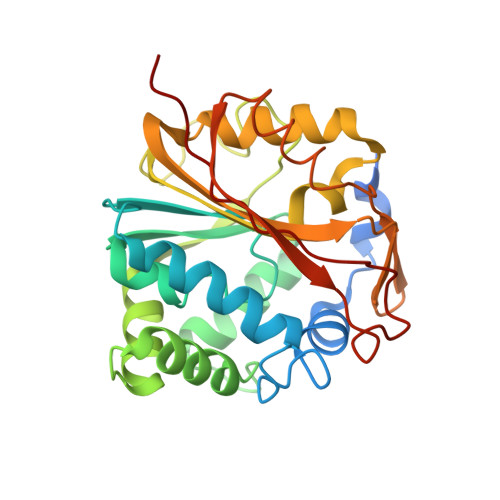
Molecular recognition of sub-micromolar inhibitors by the epinephrine-synthesizing enzyme phenylethanolamine N-methyltransferase.
McMillan, F.M., Archbold, J., McLeish, M.J., Caine, J.M., Criscione, K.R., Grunewald, G.L., Martin, J.L.(2004) J Med Chem 47: 37-44
- PubMed: 14695818 Search on PubMed
- DOI: https://doi.org/10.1021/jm0205752
- Primary Citation Related Structures:
1N7I, 1N7J - PubMed Abstract:
The crystal structures of human phenylethanolamine N-methyltransferase in complex with S-adenosyl-l-homocysteine (7, AdoHcy) and either 7-iodo-1,2,3,4-tetrahydroisoquinoline (2) or 8,9-dichloro-2,3,4,5-tetrahydro-1H-2-benzazepine (3, LY134046) were determined and compared with the structure of the enzyme complex with 7 and 7-aminosulfonyl-1,2,3,4-tetrahydroisoquinoline (1, SK&F 29661). The enzyme is able to accommodate a variety of chemically disparate functional groups on the aromatic ring of the inhibitors through adaptation of the binding pocket for this substituent and by subtle adjustments of the orientation of the inhibitors within the relatively planar binding site. In addition, the interactions formed by the amine nitrogen of all three inhibitors reinforce the hypothesis that this functional group mimics the beta-hydroxyl of norepinephrine rather than the amine. These studies provide further clues for the development of improved inhibitors for use as pharmacological probes.
- Centre for Drug Design and Development and Special Research Centre for Functional and Applied Genomics, Institute for Molecular Bioscience, University of Queensland, Brisbane QLD 4072, Australia.
Organizational Affiliation: