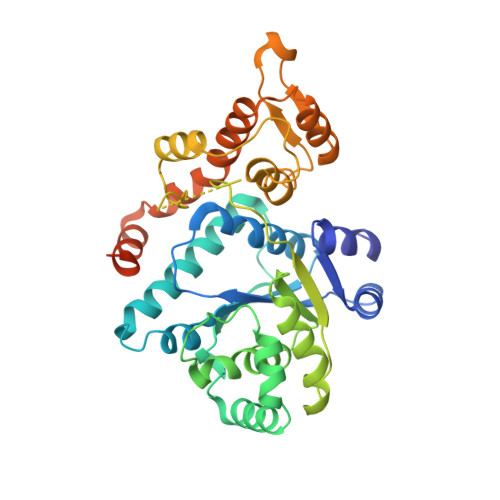
Crystal structure of a human aminoacyl-tRNA synthetase cytokine
Yang, X.-L., Skene, R.J., McRee, D.E., Schimmel, P.(2002) Proc Natl Acad Sci U S A 99: 15369-15374
- PubMed: 12427973 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.242611799
- Primary Citation Related Structures:
1N3L - PubMed Abstract:
The 20 aminoacyl-tRNA synthetases catalyze the first step of protein synthesis and establish the rules of the genetic code through aminoacylation reactions. Biological fragments of two human enzymes, tyrosyl-tRNA synthetase (TyrRS) and tryptophanyl-tRNA synthetase, connect protein synthesis to cell-signaling pathways including angiogenesis. Alternative splicing or proteolysis produces these fragments. The proangiogenic N-terminal fragment mini-TyrRS has IL-8-like cytokine activity that, like other CXC cytokines, depends on a Glu-Leu-Arg motif. Point mutations in this motif abolish cytokine activity. The full-length native TyrRS lacks cytokine activity. No structure has been available for any mammalian tRNA synthetase that, in turn, might give insight into why mini-TyrRS and not TyrRS has cytokine activities. Here, the structure of human mini-TyrRS, which contains both the catalytic and the anticodon recognition domain, is reported to a resolution of 1.18 A. The critical Glu-Leu-Arg motif is located on an internal alpha-helix of the catalytic domain, where the guanidino side chain of R is part of a hydrogen-bonding network tethering the anticodon-recognition domain back to the catalytic site. Whereas the catalytic domains of the human and bacterial enzymes superimpose, the spatial disposition of the anticodon recognition domain relative to the catalytic domain is unique in mini-TyrRS relative to the bacterial orthologs. This unique orientation of the anticodon-recognition domain can explain why the fragment mini-TyrRS, and not full-length native TyrRS, is active in cytokine-signaling pathways.
- The Skaggs Institute for Chemical Biology, BCC-379, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: