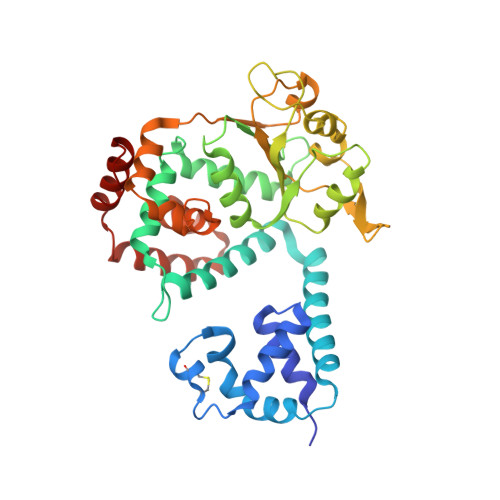
Structure of the replicative helicase of the oncoprotein SV40 large tumour antigen
Li, D., Zhao, R., Lilyestrom, W., Gai, D., Zhang, R., DeCaprio, J.A., Fanning, E., Jochimiak, A., Szakonyi, G., Chen, X.S.(2003) Nature 423: 512-518
- PubMed: 12774115 Search on PubMed
- DOI: https://doi.org/10.1038/nature01691
- Primary Citation Related Structures:
1N25 - PubMed Abstract:
The oncoprotein large tumour antigen (LTag) is encoded by the DNA tumour virus simian virus 40. LTag transforms cells and induces tumours in animals by altering the functions of tumour suppressors (including pRB and p53) and other key cellular proteins. LTag is also a molecular machine that distorts/melts the replication origin of the viral genome and unwinds duplex DNA. LTag therefore seems to be a functional homologue of the eukaryotic minichromosome maintenance (MCM) complex. Here we present the X-ray structure of a hexameric LTag with DNA helicase activity. The structure identifies the p53-binding surface and reveals the structural basis of hexamerization. The hexamer contains a long, positively charged channel with an unusually large central chamber that binds both single-stranded and double-stranded DNA. The hexamer organizes into two tiers that can potentially rotate relative to each other through connecting alpha-helices to expand/constrict the channel, producing an 'iris' effect that could be used for distorting or melting the origin and unwinding DNA at the replication fork.
- Department of Biochemistry and Molecular Genetics, University of Colorado Health Science Center, School of Medicine, Denver, Colorado 80262, USA.
Organizational Affiliation: