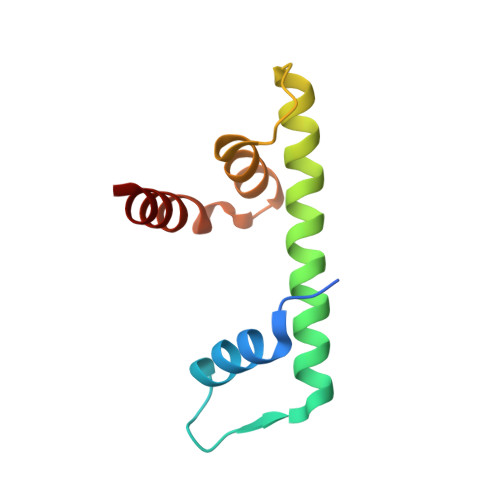
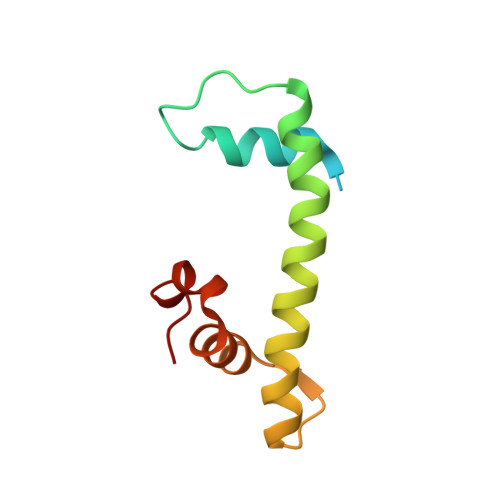
The NF-YB/NF-YC structure gives insight into DNA binding and transcription regulation by CCAAT factor NF-Y
Romier, C., Cocchiarella, F., Mantovani, R., Moras, D.(2003) J Biological Chem 278: 1336-1345
- PubMed: 12401788 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M209635200
- Primary Citation Related Structures:
1N1J - PubMed Abstract:
The heterotrimeric transcription factor NF-Y recognizes with high specificity and affinity the CCAAT regulatory element that is widely represented in promoters and enhancer regions. The CCAAT box acts in concert with neighboring elements, and its bending by NF-Y is thought to be a major mechanism required for transcription activation. We have solved the structure of the NF-YC/NF-YB subcomplex of NF-Y, which shows that the core domains of both proteins interact through histone fold motifs. This histone-like pair is closely related to the H2A/H2B and NC2alpha/NC2beta families, with features that are both common to this class of proteins and unique to NF-Y. The structure together with the modeling of the nonspecific interaction of NF-YC/NF-YB with DNA and the full NF-Y/CCAAT box complex highlight important structural features that account for different and possibly similar biological functions of the transcriptional regulators NF-Y and NC2. In particular, it emphasizes the role of the newly described alphaC helix of NF-YC, which is both important for NF-Y trimerization and a target for regulatory proteins, such as MYC and p53.
- Département de Biologie et Génomique Structurales, Institut de Génétique et de Biologie Moléculaire et Cellulaire, CNRS/INSERM/Université Louis Pasteur, 1 Rue Laurent Fries, B.P. 10142, 67404 Illkirch Cedex, France.
Organizational Affiliation: