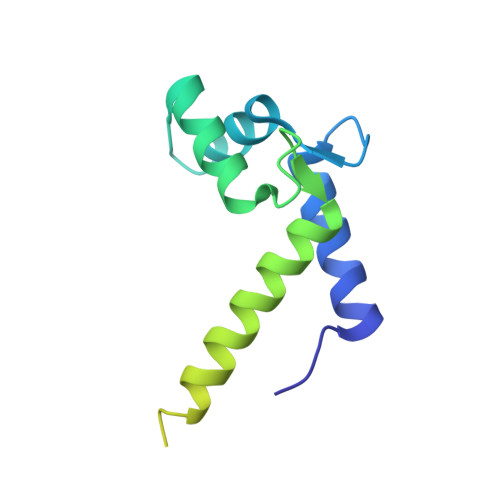
Domain flexibility in the 1.75 A resolution structure of Pb2+-calmodulin.
Wilson, M.A., Brunger, A.T.(2003) Acta Crystallogr D Biol Crystallogr 59: 1782-1792
- PubMed: 14501118 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903016846
- Primary Citation Related Structures:
1N0Y - PubMed Abstract:
Calmodulin (CaM) regulates a variety of cellular processes by interacting with a large number of proteins in a Ca(2+)-dependent manner. Conformational flexibility plays a key role in CaM function, although the full extent and detailed features of this flexibility are not fully characterized. Here, the 1.75 A resolution crystal structure of Pb(2+)-bound Paramecium tetraurelia CaM crystallized in a previously unobserved monoclinic lattice is reported. Pb(2+)-CaM is disordered in this new lattice and only a portion of each of the two molecules in the asymmetric unit can be modeled. Comparison of the structures of Ca(2+)-CaM and Pb(2+)-CaM show close agreement in the C-terminal domain but significant structural differences in the N-terminal domain. In addition, translation-libration-screw (TLS) refinement and Rosenfield difference analysis reveal inter-helical flexibility in the metal-bound N-terminal domain of the protein that is absent in the metal-bound C-terminal domain and indicates that the two structurally similar domains of CaM are dynamically distinct. These results demonstrate that TLS refinement and Rosenfield difference analysis allow detailed information about macromolecular flexibility to be extracted from X-ray diffraction data even when the crystal lattice prohibits full manifestation of this flexibility.
- Rosenstiel Basic Medical Sciences Research Center, Brandeis University, 415 South Street, Waltham, MA 02453, USA.
Organizational Affiliation: