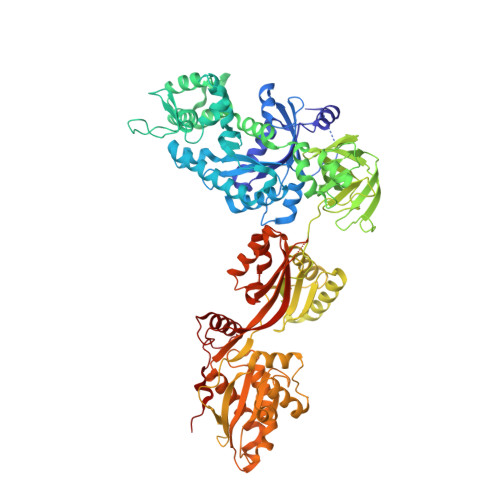
Two crystal structures demonstrate large conformational changes in the eukaryotic ribosomal translocase.
Jorgensen, R., Ortiz, P.A., Carr-Schmid, A., Nissen, P., Kinzy, T.G., Andersen, G.R.(2003) Nat Struct Biol 10: 379-385
- PubMed: 12692531 Search on PubMed
- DOI: https://doi.org/10.1038/nsb923
- Primary Citation Related Structures:
1N0U, 1N0V - PubMed Abstract:
Two crystal structures of yeast translation elongation factor 2 (eEF2) were determined: the apo form at 2.9 A resolution and eEF2 in the presence of the translocation inhibitor sordarin at 2.1 A resolution. The overall conformation of apo eEF2 is similar to that of its prokaryotic homolog elongation factor G (EF-G) in complex with GDP. Upon sordarin binding, the three tRNA-mimicking C-terminal domains undergo substantial conformational changes, while the three N-terminal domains containing the nucleotide-binding site form an almost rigid unit. The conformation of eEF2 in complex with sordarin is entirely different from known conformations observed in crystal structures of EF-G or from cryo-EM studies of EF-G-70S complexes. The domain rearrangements induced by sordarin binding and the highly ordered drug-binding site observed in the eEF2-sordarin structure provide a high-resolution structural basis for the mechanism of sordarin inhibition. The two structures also emphasize the dynamic nature of the ribosomal translocase.
- Department of Molecular Biology, Aarhus University, Gustav Wieds vej 10C, DK8000 Arhus, Denmark.
Organizational Affiliation: