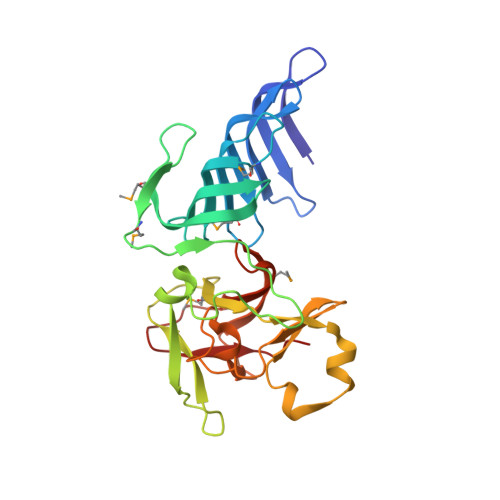
The active site of the SET domain is constructed on a knot
Jacobs, S.A., Harp, J.M., Devarakonda, S., Kim, Y., Rastinejad, F., Khorasanizadeh, S.(2002) Nat Struct Biol 9: 833-838
- PubMed: 12389038 Search on PubMed
- DOI: https://doi.org/10.1038/nsb861
- Primary Citation Related Structures:
1MT6, 1MUF - PubMed Abstract:
The SET domain contains the catalytic center of lysine methyltransferases that target the N-terminal tails of histones and regulate chromatin function. Here we report the structure of the SET7/9 protein in the absence and presence of its cofactor product, S-adenosyl-L-homocysteine (AdoHcy). A knot within the SET domain helps form the methyltransferase active site, where AdoHcy binds and lysine methylation is likely to occur. A structure-guided comparison of sequences within the SET protein family suggests that the knot substructure and active site environment are conserved features of the SET domain.
- Department of Biochemistry and Molecular Genetics, University of Virginia Health System, Charlottesville, Virginia 22908, USA.
Organizational Affiliation: