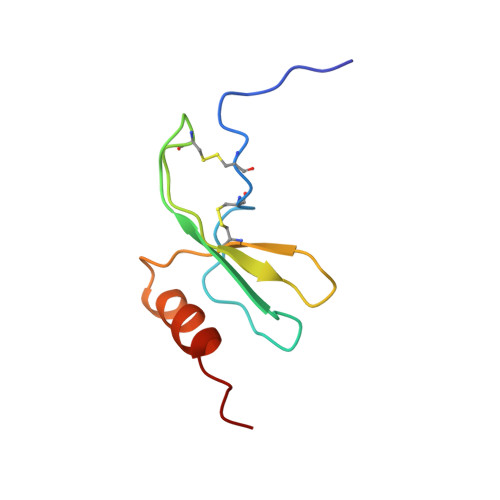
Solution structure of GRO/melanoma growth stimulatory activity determined by 1H NMR spectroscopy.
Kim, K.S., Clark-Lewis, I., Sykes, B.D.(1994) J Biological Chem 269: 32909-32915
- PubMed: 7806518 Search on PubMed
- Primary Citation Related Structures:
1MSG, 1MSH - PubMed Abstract:
The three-dimensional solution structure of the growth-related protein-alpha/melanoma growth stimulatory activity (GRO/MGSA) has been solved by two-dimensional 1H nuclear magnetic resonance spectroscopy. The GRO/MGSA monomer consists of an NH2-terminal loop, a three-stranded antiparallel beta-sheet, and a COOH-terminal alpha-helix. Dimerization, which is apparent under the experimental conditions used (2 mM, pH 5.10, 30 degrees C), results in a six-stranded antiparallel beta-sheet and a pair of helices with 2-fold symmetry. While the basic fold is similar to that seen for interleukin-8 (IL-8) (Clore, G. M., Appella, E., Yamada, M., Matsushima, K., and Gronenborn, A. M. (1990) Biochemistry, 29, 1689-1696), there are differences in the ELR motif (residues 6-8), the turn involving residues 31-36, which is linked to the NH2-terminal region through the 9-35 disulfide bond. The most significant differences are in the NH2-terminal loop (residues 12-23). In IL-8, all the corresponding regions have been shown to be required for receptor binding (Clark-Lewis, I., Dewald, B., Loetscher, M., Moser, B., and Baggiolini, M. (1994) J. Biol. Chem. 269, 16075-16081). The structural differences thus have been identified between GRO/MGSA and IL-8 could contribute to their different receptor binding specificities.
- Protein Engineering Network of Centres of Excellence (PENCE), University of Alberta, Edmonton, Canada.
Organizational Affiliation: