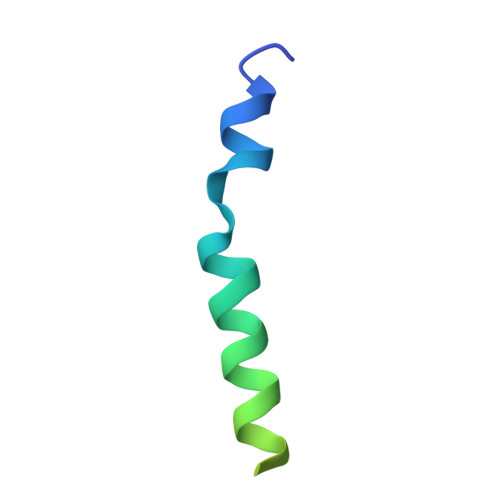
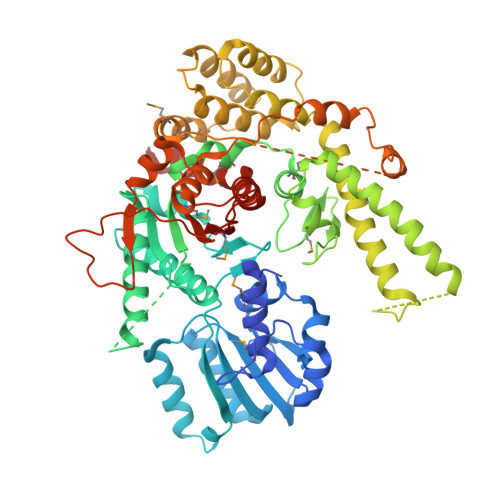
Structural basis for the Golgi membrane recruitment of Sly1p by Sed5p
Bracher, A., Weissenhorn, W.(2002) EMBO J 21: 6114-6124
- PubMed: 12426383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdf608
- Primary Citation Related Structures:
1MQS - PubMed Abstract:
Cytosolic Sec1/munc18-like proteins (SM proteins) are recruited to membrane fusion sites by interaction with syntaxin-type SNARE proteins, constituting indispensable positive regulators of intracellular membrane fusion. Here we present the crystal structure of the yeast SM protein Sly1p in complex with a short N-terminal peptide derived from the Golgi-resident syntaxin Sed5p. Sly1p folds, similarly to neuronal Sec1, into a three-domain arch-shaped assembly, and Sed5p interacts in a helical conformation predominantly with domain I of Sly1p on the opposite site of the nSec1/syntaxin-1-binding site. Sequence conservation of the major interactions suggests that homologues of Sly1p as well as the paralogous Vps45p group bind their respective syntaxins in the same way. Furthermore, we present indirect evidence that nSec1 might be able to contact syntaxin 1 in a similar fashion. The observed Sly1p-Sed5p interaction mode therefore indicates how SM proteins can stay associated with the assembling fusion machinery in order to participate in late fusion steps.
- European Molecular Biology Laboratory, 6 rue Jules Horowitz, 38042 Grenoble, France. bracher@embl-grenoble.fr
Organizational Affiliation: