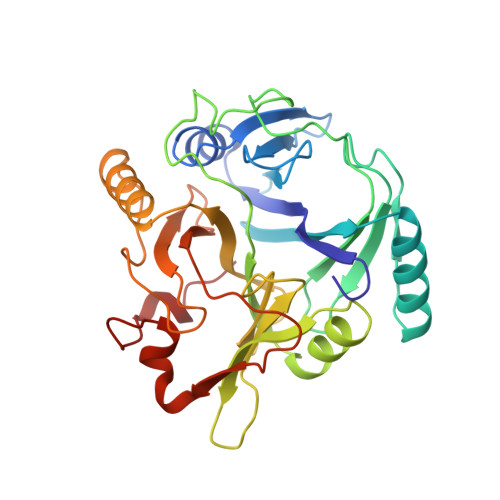
An archetypical extradiol-cleaving catecholic dioxygenase: the crystal structure of catechol 2,3-dioxygenase (metapyrocatechase) from Ppseudomonas putida mt-2.
Kita, A., Kita, S., Fujisawa, I., Inaka, K., Ishida, T., Horiike, K., Nozaki, M., Miki, K.(1999) Structure 7: 25-34
- PubMed: 10368270 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(99)80006-9
- Primary Citation Related Structures:
1MPY - PubMed Abstract:
Catechol dioxygenases catalyze the ring cleavage of catechol and its derivatives in either an intradiol or extradiol manner. These enzymes have a key role in the degradation of aromatic molecules in the environment by soil bacteria. Catechol 2, 3-dioxygenase catalyzes the incorporation of dioxygen into catechol and the extradiol ring cleavage to form 2-hydroxymuconate semialdehyde. Catechol 2,3-dioxygenase (metapyrocatechase, MPC) from Pseudomonas putida mt-2 was the first extradiol dioxygenase to be obtained in a pure form and has been studied extensively. The lack of an MPC structure has hampered the understanding of the general mechanism of extradiol dioxygenases. The three-dimensional structure of MPC has been determined at 2.8 A resolution by the multiple isomorphous replacement method. The enzyme is a homotetramer with each subunit folded into two similar domains. The structure of the MPC subunit resembles that of 2,3-dihydroxybiphenyl 1,2-dioxygenase, although there is low amino acid sequence identity between these enzymes. The active-site structure reveals a distorted tetrahedral Fe(II) site with three endogenous ligands (His153, His214 and Glu265), and an additional molecule that is most probably acetone. The present structure of MPC, combined with those of two 2,3-dihydroxybiphenyl 1,2-dioxygenases, reveals a conserved core region of the active site comprising three Fe(II) ligands (His153, His214 and Glu265), one tyrosine (Tyr255) and two histidine (His199 and His246) residues. The results suggest that extradiol dioxygenases employ a common mechanism to recognize the catechol ring moiety of various substrates and to activate dioxygen. One of the conserved histidine residues (His199) seems to have important roles in the catalytic cycle.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: