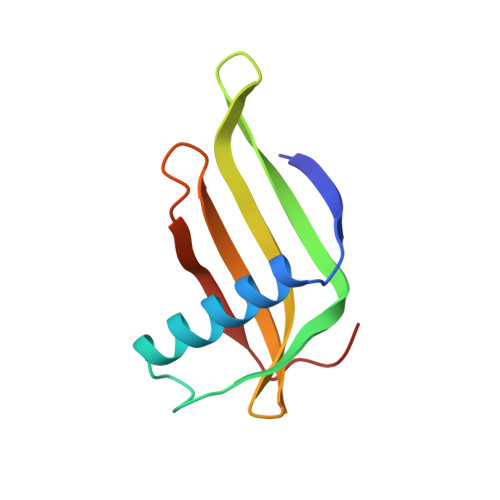
Two crystal structures of a potently sweet protein. Natural monellin at 2.75 A resolution and single-chain monellin at 1.7 A resolution.
Somoza, J.R., Jiang, F., Tong, L., Kang, C.H., Cho, J.M., Kim, S.H.(1993) J Mol Biology 234: 390-404
- PubMed: 8230222
- DOI: https://doi.org/10.1006/jmbi.1993.1594
- Primary Citation Related Structures:
1MOL - PubMed Abstract:
Two refined structures of the sweet-tasting protein monellin are presented. The structure of natural monellin has been refined at 2.75 A resolution. The final model consists of four monellin molecules in the asymmetric unit, encompassing 3136 non-hydrogen atoms. The crystallographic R-factor is 0.193 for the 8853 reflections between 6.0 A and 2.75 A resolution, and the root-mean-square deviations from ideality are 0.017 A for bond lengths and 3.6 degrees for bond angles. The refined structure generally confirms, with some difference in detail, the initial backbone model of monellin that was based on 3.0 A resolution data. Single-chain monellin (scm) was genetically engineered by fusing the two chains of monellin into a single 94-residue polypeptide. Using the refined monellin coordinates as a search model, the crystal structure of scm has been solved with the techniques of molecular replacement, and has been refined against data to 1.7 A resolution. The final model consists of two scm molecules per asymmetric unit, and includes 137 bound water molecules. The crystallographic R-factor for this model is 0.174 for the 15,053 reflections (magnitude of FO magnitude of > 2 sigma (FO)) between 6.0 A and 1.7 A resolution. The root-mean-square deviations from ideal bond lengths and angles are 0.015 A and 2.86 degrees, respectively, and the average coordinate error is approximately 0.2 A, as estimated from a Luzzati plot. The error in the model was also estimated by comparing the two molecules in the asymmetric unit. The most significant differences between the two molecules occur in loop regions and at the C terminus of the protein, and are generally correlated to differences in crystal packing contacts. Linking the two chains of monellin has not substantially altered the structure beyond the region immediately surrounding the new peptide bond. Like natural monellin, the conformation of scm is dominated by a 17-residue alpha-helix folded into the concave side of a twisted, five-strand anti-parallel beta-sheet. We expect that the availability of a high-resolution structure of scm, along with the convenience of producing site-specific mutants of this protein, will make scm a good model with which to probe the structural basis of sweetness.
- Graduate Group in Biophysics, University of California at Berkeley 94720.
Organizational Affiliation: