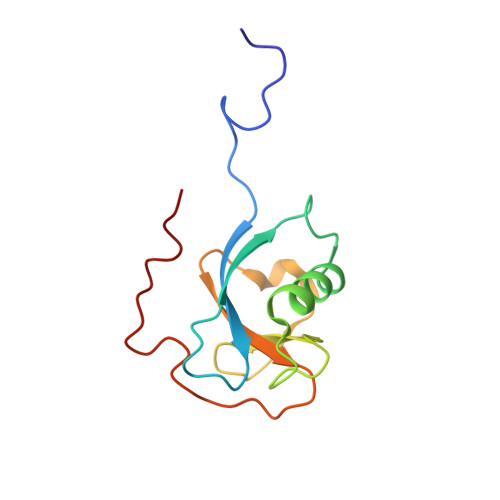
The DCX-domain Tandems of Doublecortin and Doublecortin-like Kinase
Kim, M.H., Cierpicki, T., Derewenda, U., Krowarsch, D., Feng, Y., Devedjiev, Y., Dauter, Z., Walsh, C.A., Otlewski, J., Bushweller, J.H., Derewenda, Z.S.(2003) Nat Struct Biol 10: 324-333
- PubMed: 12692530 Search on PubMed
- DOI: https://doi.org/10.1038/nsb918
- Primary Citation Related Structures:
1MFW, 1MG4, 1MJD - PubMed Abstract:
The doublecortin-like domains (DCX), which typically occur in tandem, are novel microtubule-binding modules. DCX tandems are found in doublecortin, a 360-residue protein expressed in migrating neurons; the doublecortin-like kinase (DCLK); the product of the RP1 gene that is responsible for a form of inherited blindness; and several other proteins. Mutations in the gene encoding doublecortin cause lissencephaly in males and the 'double-cortex syndrome' in females. We here report a solution structure of the N-terminal DCX domain of human doublecortin and a 1.5 A resolution crystal structure of the equivalent domain from human DCLK. Both show a stable, ubiquitin-like tertiary fold with distinct structural similarities to GTPase-binding domains. We also show that the C-terminal DCX domains of both proteins are only partially folded. In functional assays, the N-terminal DCX domain of doublecortin binds only to assembled microtubules, whereas the C-terminal domain binds to both microtubules and unpolymerized tubulin.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, Virginia 22908-0736, USA.
Organizational Affiliation: