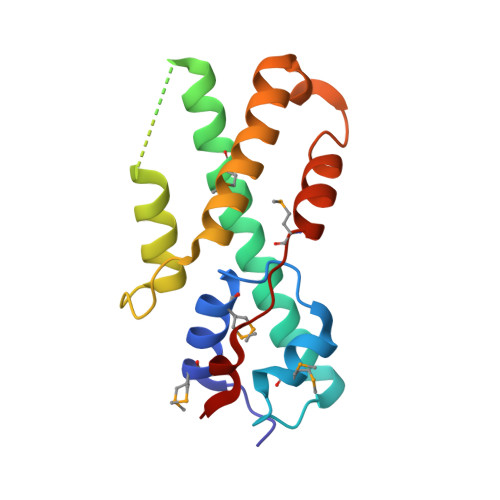
Structure of the DNA Binding Region of Prospero Reveals a Novel Homeo-Prospero Domain
Ryter, J.M., Doe, C.Q., Matthews, B.W.(2002) Structure 10: 1541-1549
- PubMed: 12429095 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00883-3
- Primary Citation Related Structures:
1MIJ - PubMed Abstract:
The Prospero transcription factor promotes neural differentiation in Drosophila, and its activity is tightly regulated by modulating its subcellular localization. Prospero is exported from the nucleus of neural precursors but imported into the nucleus of daughter cells, which is necessary for their proper differentiation. Prospero has a highly divergent putative homeodomain adjacent to a conserved Prospero domain; both are required for sequence-specific DNA binding. Here we show that the structure of these two regions consists of a single structural unit (a homeo-prospero domain), in which the Prospero domain region is in position to contribute to DNA binding and also to mask a defined nuclear export signal that is within the putative homeodomain region. We propose that the homeo-prospero domain coordinately regulates Prospero nuclear localization and DNA binding specificity.
- Institute of Molecular Biology, Howard Hughes Medical Institute and Department of Physics, 1229 University of Oregon, Eugene, OR 97403, USA.
Organizational Affiliation: