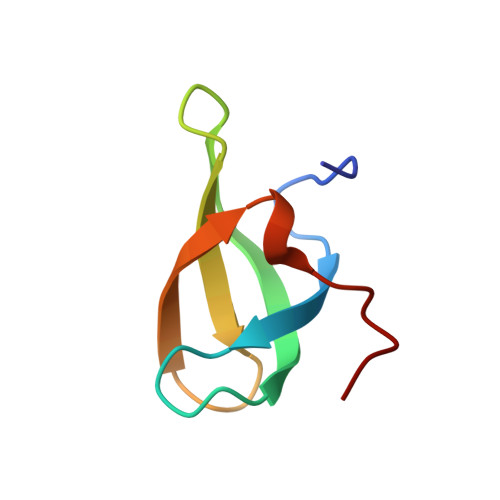
High Resolution X-ray and NMR Structures of the SMN Tudor Domain: conformational variation in the binding site for symmetrically dimethylated arginine residues
Sprangers, R., Groves, M.R., Sinning, I., Sattler, M.(2003) J Mol Biology 327: 507-520
- PubMed: 12628254 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00148-7
- Primary Citation Related Structures:
1MHN - PubMed Abstract:
The SMN protein, which is linked to spinal muscular atrophy (SMA), plays an important role in the assembly of the spliceosomal small nuclear ribonucleoprotein complexes. This function requires binding of SMN to the arginine-glycine (RG) rich C-terminal tails of the Sm proteins, which contain symmetrically dimethylated arginine residues (sDMA) in vivo. Using NMR titrations, we show that the SMN Tudor domain recognizes these sDMAs in the methylated RG repeats. Upon complex formation a cluster of conserved aromatic residues in the SMN Tudor domain interacts with the sDMA methyl groups. We present two high resolution structures of the uncomplexed SMN Tudor domain, a 1.8A crystal structure and an NMR structure that has been refined against a large number of backbone and side-chain residual dipolar couplings. The backbone conformation of both structures is very similar, however, differences are observed for the cluster of conserved aromatic side-chains in the sDMA binding pocket. In order to validate these variations we introduce a novel application of residual dipolar couplings for aromatic rings. We show that structural information can be derived from aromatic ring residual dipolar couplings, even in the presence of internal motions such as ring flipping. These residual dipolar couplings and ring current shifts independently confirm that the SMN Tudor domain adopts two different conformations in the sDMA binding pocket. The observed structural variations may play a role for the recognition of sDMAs.
- Structural and Computational Biology Programme, EMBL Heidelberg, Meyerhofstr. 1, Germany. spranger@embl.de
Organizational Affiliation: