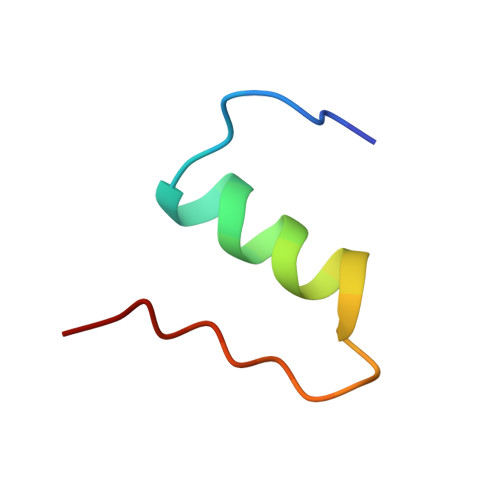
Three-dimensional solution structure of an insulin dimer. A study of the B9(Asp) mutant of human insulin using nuclear magnetic resonance, distance geometry and restrained molecular dynamics.
Jorgensen, A.M., Kristensen, S.M., Led, J.J., Balschmidt, P.(1992) J Mol Biology 227: 1146-1163
- PubMed: 1433291 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90527-q
- Primary Citation Related Structures:
1MHI - PubMed Abstract:
The solution structure of the B9(Asp) mutant of human insulin has been determined by two-dimensional 1H nuclear magnetic resonance spectroscopy. Thirty structures were calculated by distance geometry from 451 interproton distance restraints based on intra-residue, sequential and long-range nuclear Overhauser enhancement data, 17 restraints on phi torsional angles obtained from 3JH alpha HN coupling constants, and the restraints from 17 hydrogen bonds, and the three disulphide bridges. The distance geometry structures were optimized using restrained molecular dynamics (RMD) and energy minimization. The average root-mean-square deviation for the best 20 RMD refined structures is 2.26 A for the backbone and 3.14 A for all atoms if the less well-defined N and C-terminal residues are excluded. The helical regions are better defined, with root-mean-square deviation values of 1.11 A for the backbone and 2.03 A for all atoms. The data analysis and the calculations show that B9(Asp) insulin, in water solution at the applied pH (1.8 to 1.9), is a well-defined dimer with no detectable difference between the two monomers. The association of the two monomers in the solution dimer is relatively loose as compared with the crystal dimer. The overall secondary and tertiary structures of the monomers in the 2Zn crystal hexamer is found to be preserved. The conformation-averaged NMR structures obtained for the monomer is close to the structure of molecule 1 in the hexamer of the 2Zn insulin crystal. However, minor, but significant deviations from this structure, as well as from the structure of monomeric insulin in solution, exist and are ascribed to the absence of the hexamer and crystal packing forces, and to the presence of monomer-monomer interactions, respectively. Thus, the monomer in the solution dimer shows a conformation similar to that of the crystal monomer in molecular regions close to the monomer-monomer interface, whereas it assumes a conformation similar to that of the solution structure of monomeric insulin in other regions, suggesting that B9(Asp) insulin adopts a monomer-like conformation when this is not inconsistent with the monomer-monomer arrangement in the dimer.
- Department of Chemistry, University of Copenhagen, H.C. Orsted Institute, Denmark.
Organizational Affiliation: