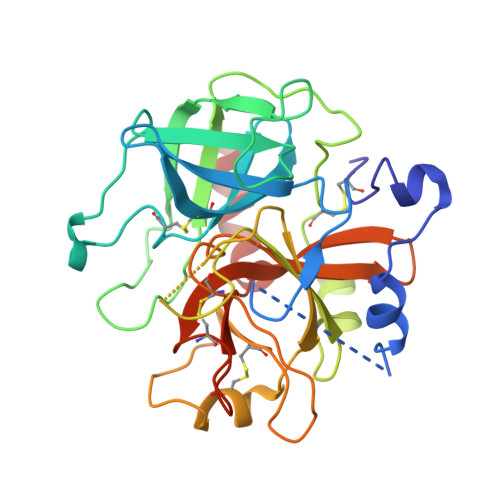
Crystal structure of the anticoagulant slow form of thrombin
Pineda, A.O., Savvides, S., Waksman, G., Di Cera, E.(2002) J Biological Chem 277: 40177-40180
- PubMed: 12205081 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C200465200
- Primary Citation Related Structures:
1MH0 - PubMed Abstract:
Using the thrombin mutant R77aA devoid of the site of autoproteolytic degradation at exosite I, we have solved for the first time the structure of thrombin free of any inhibitors and effector molecules and stabilized in the Na(+)-free slow form. The slow form shows subtle differences compared with the currently available structures of the Na(+)-bound fast form that carry inhibitors at the active site or exosite I. The most notable differences are the displacement of Asp-189 in the S1 specificity pocket, a downward shift of the 190-193 strand, a rearrangement of the side chain of Glu-192, and a significant shift in the position of the catalytic Ser-195 that is no longer within H-bonding distance from His-57. The structure of the slow form explains the reduced specificity toward synthetic and natural substrates and suggests a molecular basis for its anticoagulant properties.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, MO 63110, USA.
Organizational Affiliation: