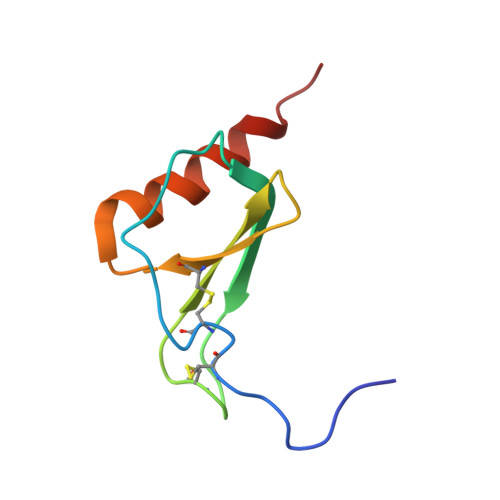
The solution structure of melanoma growth stimulating activity.
Fairbrother, W.J., Reilly, D., Colby, T.J., Hesselgesser, J., Horuk, R.(1994) J Mol Biology 242: 252-270
- PubMed: 8089846
- DOI: https://doi.org/10.1006/jmbi.1994.1577
- Primary Citation Related Structures:
1MGS - PubMed Abstract:
The solution structure of melanoma growth stimulating activity (MGSA), a dimeric chemokine consisting of 73 residues per monomer, has been determined using two-dimensional homonuclear and three-dimensional heteronuclear NMR spectroscopy. Structure calculations were carried out using a hybrid distance geometry-simulated annealing approach with the programs DGII and X-PLOR. The structure is based on a total of 2362 experimental restraints, comprising 2150 NOE-derived distance restraints (2076 unambiguous intrasubunit restraints, 60 unambiguous intersubunit restraints, and 14 ambiguous restraints with potential contributions from both intra- and intersubunit NOEs), 84 distance restraints for 42 backbone hydrogen bonds, and 128 torsion angle restraints. The ambiguous distance restraints were treated using a target function which accounts for both intra- and intermolecular contributions to the NOE intensity. A total of 25 structures were calculated, with the backbone (N, C alpha, C) atomic r.m.s. distribution about the mean coordinates for residues 8 to 69 being 0.44(+/- 0.10) A for the dimer and 0.34(+/- 0.07) A for the individual monomers. The N- and C-terminal residues (1 to 7 and 70 to 73, respectively) are disordered. The overall structure of the MGSA dimer is similar to that reported previously for the NMR and X-ray structures of interleukin-8 (IL-8), and consists of a six-stranded antiparallel beta-sheet packed against two C-terminal antiparallel alpha-helices. A best fit superposition of the NMR structure of MGSA on the X-ray and NMR structures of IL-8 yields backbone atomic r.m.s. differences of 0.99 and 1.28 A, respectively for individual monomers, and 1.08 and 1.82 A, respectively for the dimers (using MGSA residues 8 to 14 and 19 to 69). In general, the MGSA structure resembles the IL-8 X-ray structure more than it does the IL-8 NMR structure. At the tertiary (monomer) level the two main differences between the MGSA solution structure and IL-8 NMR structure involve the loops between residues 14 to 19 and between residues 30 to 38. At the quaternary (dimer) level the difference results from differing angles between the beta-strands which form the dimer interface, and is manifest as a different interhelical separation (distance of closest approach between the two helices is 15.3 A in the IL-8 NMR structure and 11.7 (+/- 0.4) A in the MGSA structure).
- Department of Protein Engineering, Genentech, Inc., South San Francisco, CA 94080-4990.
Organizational Affiliation: