Induced structural changes of 7SL RNA during the assembly of human signal recognition particle
Kuglstatter, A., Oubridge, C., Nagai, K.(2002) Nat Struct Biol 9: 740-744
- PubMed: 12244299 Search on PubMed
- DOI: https://doi.org/10.1038/nsb843
- Primary Citation Related Structures:
1MFQ - PubMed Abstract:
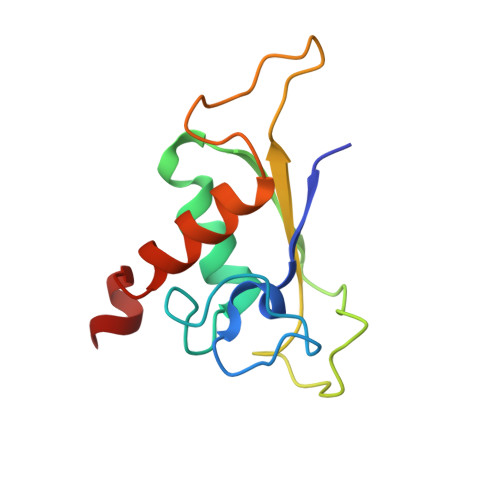
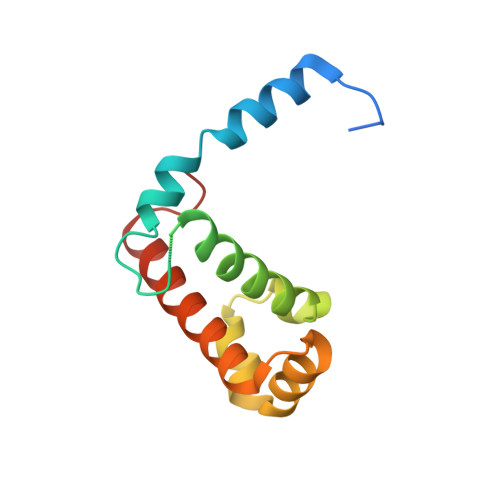
The eukaryotic signal recognition particle (SRP) is a cytoplasmic ribonucleoprotein particle that targets secretory and membrane proteins to the endoplasmic reticulum. The binding of SRP54 to the S domain of 7SL RNA is highly dependent on SRP19. Here we present the crystal structure of a human SRP ternary complex consisting of SRP19, the M domain of SRP54 and the S domain of 7SL RNA. Upon binding of the M domain of SRP54 to the 7SL RNA-SRP19 complex, the asymmetric loop of helix 8 in 7SL RNA collapses. The bases of the four nucleotides in the long strand of the asymmetric loop continuously stack and interact with the M domain, whereas the two adenines in the short strand flip out and form two A-minor motifs with helix 6. This stabilizing interaction is only possible when helix 6 has been positioned parallel to helix 8 by the prior binding of SRP19 to the tetraloops of helices 6 and 8. Hence, the crystal structure of the ternary complex suggests why SRP19 is necessary for the stable binding of SRP54 to the S domain RNA.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: