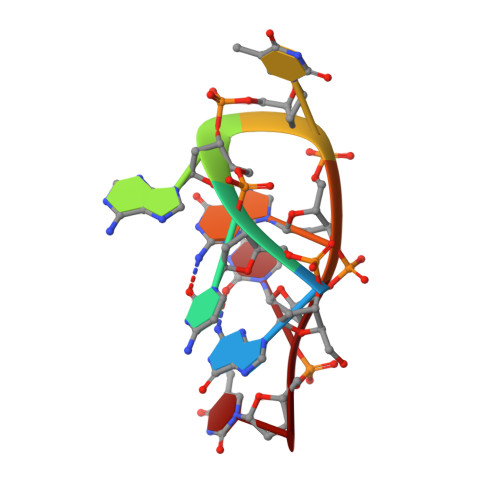
Crystal structure of the complementary quadruplex formed by d(GCATGCT) at atomic resolution
Thorpe, J.H., Teixeira, S.C.M., Gale, B.C., Cardin, C.J.(2003) Nucleic Acids Res 31: 844-849
- PubMed: 12560479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkg168
- Primary Citation Related Structures:
1MF5 - PubMed Abstract:
Here we report the crystal structure of the DNA heptanucleotide sequence d(GCATGCT) determined to a resolution of 1.1 A. The sequence folds into a complementary loop structure generating several unusual base pairings and is stabilised through cobalt hexammine and highly defined water sites. The single stranded loop is bound together through the G(N2)-C(O2) intra-strand H-bonds for the available G/C residues, which form further Watson-Crick pairings to a complementary sequence, through 2-fold symmetry, generating a pair of non-planar quadruplexes at the heart of the structure. Further, four adenine residues stack in pairs at one end, H-bonding through their N7-N6 positions, and are additionally stabilised through two highly conserved water positions at the structural terminus. This conformation is achieved through the rotation of the central thymine base at the pinnacle of the loop structure, where it stacks with an adjacent thymine residue within the lattice. The crystal packing yields two halved biological units, each related across a 2-fold symmetry axis spanning a cobalt hexammine residue between them, which stabilises the quadruplex structure through H-bonds to the phosphate oxygens and localised hydration.
- The University of Reading, School of Chemistry, Whiteknights, Reading, Berkshire RG6 6AD, UK.
Organizational Affiliation: