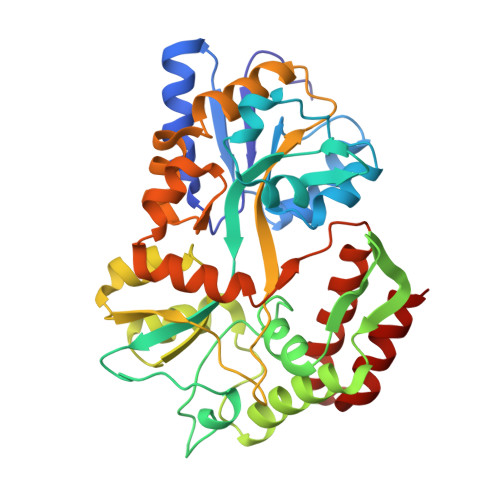
Refined structures of two insertion/deletion mutants probe function of the maltodextrin binding protein.
Sharff, A.J., Rodseth, L.E., Szmelcman, S., Hofnung, M., Quiocho, F.A.(1995) J Mol Biology 246: 8-13
- PubMed: 7853407
- DOI: https://doi.org/10.1006/jmbi.1994.0059
- Primary Citation Related Structures:
1MDP, 1MDQ - PubMed Abstract:
The X-ray structures of the maltose bound forms of two insertion/deletion mutants of the Escherichia coli maltodextrin binding protein, MalE322 and MalE178, have been determined and refined. MalE322 involves a one residue deletion, two residue insertion in a hinge segment connecting the two (N and C) domains of the protein, an area already identified as being critical for the correct functioning of the protein. MalE178 involves a nine residue deletion and two residue insertion in a helix at the periphery of the C-domain. The function of both mutant proteins is similar to the wild-type, although MalE322 increases the ability to transport maltose and maltodextrin whilst inhibiting the ability of the cell to grow on dextrins. Both proteins exhibit very localized and conservative conformational changes due to their mutations. The structure of MalE322 shows some deformation of the third hinge strand, indicating the likely cause of change in its biochemistry. MalE178 is stable and its activity virtually unchanged from the wild-type. This is most likely due to the long distance of the mutation from the binding site and conservation of the number of interactions between the area around the deletion site and the main body of the protein.
- Howard Hughes Medical Institute, Baylor College of Medicine, Houston, TX 77030.
Organizational Affiliation: