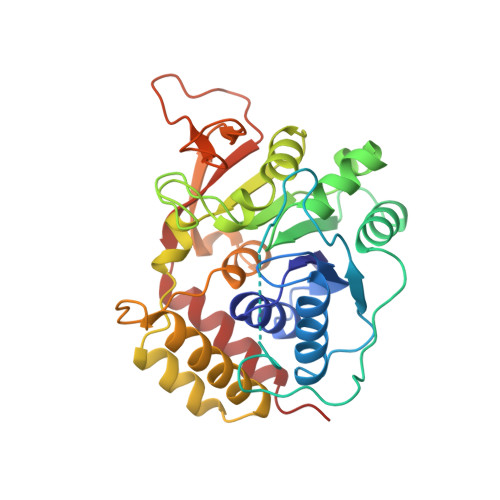
Three-dimensional structure of the inosine-uridine nucleoside N-ribohydrolase from Crithidia fasciculata.
Degano, M., Gopaul, D.N., Scapin, G., Schramm, V.L., Sacchettini, J.C.(1996) Biochemistry 35: 5971-5981
- PubMed: 8634238 Search on PubMed
- DOI: https://doi.org/10.1021/bi952999m
- Primary Citation Related Structures:
1MAS - PubMed Abstract:
Protozoan parasites rely on the host for purines since they lack a de novo synthetic pathway. Crithidia fasciculata salvages exogenous inosine primarily through hydrolysis of the N-ribosidic bond using several nucleoside hydrolases. The most abundant nucleoside hydrolase is relatively nonspecific but prefers inosine and uridine as substrates. Here we report the three-dimensional structure of the inosine-uridine nucleoside hydrolase (IU-NH) from C. fasciculata determined by X-ray crystallography at a nominal resolution of 2.5 A. The enzyme has an open (alpha, beta) structure which differs from the classical dinucleotide binding fold. IU-nucleoside hydrolase is composed of a mixed eight-stranded beta sheet surrounded by six alpha helices and a small C-terminal lobe composed of four alpha helices. Two short antiparallel beta strands are involved in intermolecular contacts. The catalytic pocket is located at the C-terminal end of beta strands beta 1 and beta 4. Four aspartate residues are located at the bottom of the cavity in a geometry which suggests interaction with the ribose moiety of the nucleoside. These groups could provide the catalytically important interactions to the ribosyl hydroxyls and the stabilizing anion for the oxycarbonium-like transition state. Histidine 241, located on the side of the active site cavity, is the proposed proton donor which facilitates purine base departure [Gopaul, D. N., Meyer, S. L., Degano, M., Sacchettini, J. C., & Schramm, V. L. (1996) Biochemistry 35, 5963-5970]. The substrate binding site is unlike that from purine nucleoside phosphorylase, phosphoribosyltransferases, or uracil DNA glycosylase and thus represents a novel architecture for general acid-base catalysis. This detailed knowledge of the architecture of the active site, together with the previous transition state analysis [Horenstein, B. A., Parkin, D. W., Estupiñán, B., & Schramm, V. L. (1991) Biochemistry 30, 10788-10795], allows analysis of the interactions leading to catalysis and an explanation for the tight-binding inhibitors of the enzyme [Schramm, V. L., Horenstein, B. A., & Kline, P. C. (1994) J. Biol. Chem. 269, 18259-18262].
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461, USA.
Organizational Affiliation: