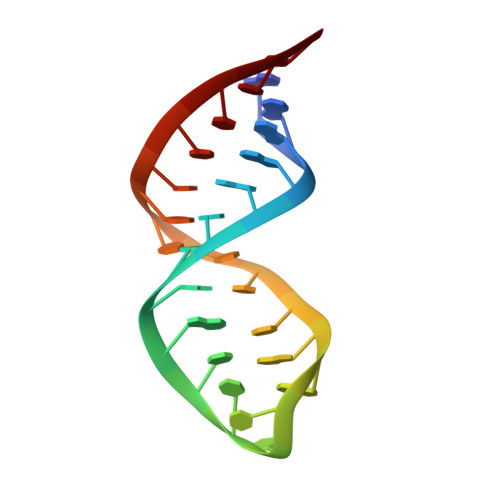
Solution structure of the influenza A virus cRNA promoter: implications for differential recognition of viral promoter structures by RNA-dependent RNA polymerase
Park, C.-J., Bae, S.-H., Lee, M.-K., Varani, G., Choi, B.-S.(2003) Nucleic Acids Res 31: 2824-2832
- PubMed: 12771209 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkg387
- Primary Citation Related Structures:
1M82 - PubMed Abstract:
Influenza A virus replication requires the interaction of viral RNA-dependent RNA polymerase (RdRp) with promoters in both the RNA genome (vRNA) and the full-length complementary RNA (cRNA) which serve as templates for the generation of new vRNAs. Although RdRp binds both promoters effectively, it must also discriminate between them because they serve different functional roles in the viral life cycle. Even though the inherent asymmetry between two RNA promoters is considered as a cause of the differential recognition by the RdRp, the structural basis for the ability of the RdRp to recognize the RNA promoters and discriminate effectively between them remains unsolved. Here we report the structure of the cRNA promoter of influenza A virus as determined by heteronuclear magnetic resonance spectroscopy. The terminal region is extremely unstable and does not have a rigid structure. The major groove of the internal loop is widened by the displacement of a novel A*(UU) motif toward the minor groove. These internal loop residues show distinguishable dynamic characters, with differing motional timescales for each residue. Comparison of the cRNA promoter structure with that of the vRNA promoter reveals common structural and dynamic elements in the internal loop, but also differences that provide insight into how the viral RdRp differentially recognizes the cRNA and vRNA promoters.
- Department of Chemistry and National Creative Research Initiative Center, Korea Advanced Institute of Science and Technology, 373-1 Guseong-dong, Yuseong-gu, Daejon 305-701, Korea.
Organizational Affiliation: