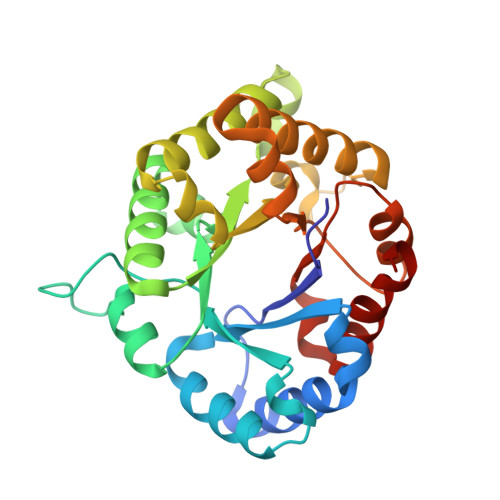
Structure and Inactivation of Triosephosphate Isomerase from Entamoeba histolytica
Rodriguez-Romero, A., Hernandez-Santoyo, A., Del Pozo-Yauner, L., Kornhauser, A., Fernandez-Velasco, D.A.(2002) J Mol Biology 322: 669-675
- PubMed: 12270704 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00809-4
- Primary Citation Related Structures:
1M6J - PubMed Abstract:
Triosephosphate isomerase (TIM) has been proposed as a target for drug design. TIMs from several parasites have a cysteine residue at the dimer interface, whose derivatization with thiol-specific reagents induces enzyme inactivation and aggregation. TIMs lacking this residue, such as human TIM, are less affected. TIM from Entamoeba histolytica (EhTIM) has the interface cysteine residue and presents more than ten insertions when compared with the enzyme from other pathogens. To gain further insight into the role that interface residues play in the stability and reactivity of these enzymes, we determined the high-resolution structure and characterized the effect of methylmethane thiosulfonate (MMTS) on the activity and conformational properties of EhTIM. The structure of this enzyme was determined at 1.5A resolution using molecular replacement, observing that the dimer is not symmetric. EhTIM is completely inactivated by MMTS, and dissociated into stable monomers that possess considerable secondary structure. Structural and spectroscopic analysis of EhTIM and comparison with TIMs from other pathogens reveal that conformational rearrangements of the interface after dissociation, as well as intramonomeric contacts formed by the inserted residues, may contribute to the unusual stability of the derivatized EhTIM monomer.
- Laboratorio Universitario de Estructura de Proteínas and Departamento de Bioquímica, Instituto de Química, Universidad Nacional Autónoma de México, DF, Mexico. adela@servidor.unam.mx
Organizational Affiliation: