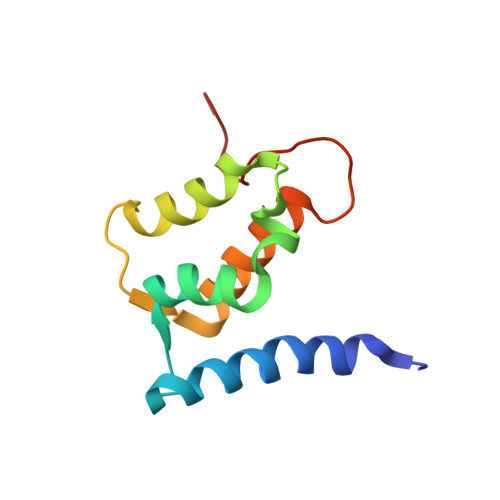
Solution structure of human Mts1 (S100A4) as determined by NMR spectroscopy.
Vallely, K.M., Rustandi, R.R., Ellis, K.C., Varlamova, O., Bresnick, A.R., Weber, D.J.(2002) Biochemistry 41: 12670-12680
- PubMed: 12379109 Search on PubMed
- DOI: https://doi.org/10.1021/bi020365r
- Primary Citation Related Structures:
1M31 - PubMed Abstract:
Mts1 is a member of the S100 family of Ca2+-binding proteins and is implicated in promoting tumor progression and metastasis. To better understand the structure-function relationships of this protein and to begin characterizing its Ca2+-dependent interaction with protein binding targets, the three-dimensional structure of mts1 was determined in the apo state by NMR spectroscopy. As with other S100 protein family members, mts1 is a symmetric homodimer held together by noncovalent interactions between two helices from each subunit (helices 1, 4, 1', and 4') to form an X-type four-helix bundle. Each subunit of mts1 has two EF-hand Ca2+-binding domains: a pseudo-EF-hand (or S100-hand) and a typical EF-hand that are brought into proximity by a small two-stranded antiparallel beta-sheet. The S100-hand is formed by helices 1 and 2, and is similar in conformation to other members of the S100 family. In the typical EF-hand, the position of helix 3 is similar to that of another member of the S100 protein family, calcyclin (S100A6), and less like that of other S100 family members for which three-dimensional structures are available in the calcium-free state (e.g., S100B and S100A1). The differences in the position of helix 3 in the apo state of these four S100 proteins are likely due to variations in the amino acid sequence in the C-terminus of helix 4 and in loop 2 (the hinge region) and could potentially be used to subclassify the S100 protein family.
- Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, 108 North Greene Street, Baltimore, Maryland 21201, USA.
Organizational Affiliation: