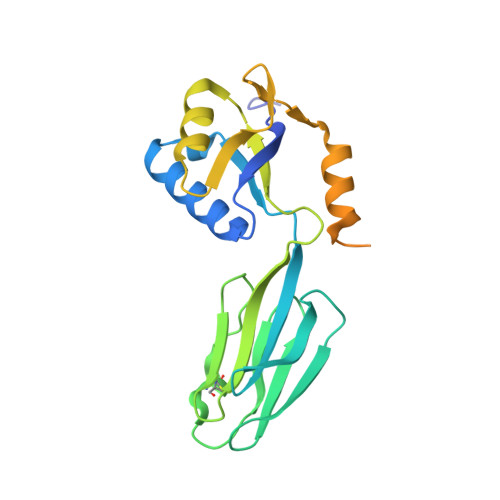
Crystal structures of transcription factor NusG in light of its nucleic acid- and protein-binding activities
Steiner, T., Kaiser, J.T., Marinkovic, S., Huber, R., Wahl, M.C.(2002) EMBO J 21: 4641-4653
- PubMed: 12198166 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdf455
- Primary Citation Related Structures:
1M1G, 1M1H - PubMed Abstract:
Microbial transcription modulator NusG interacts with RNA polymerase and termination factor rho, displaying striking functional homology to eukaryotic Spt5. The protein is also a translational regulator. We have determined crystal structures of Aquifex aeolicus NusG showing a modular design: an N-terminal RNP-like domain, a C-terminal element with a KOW sequence motif and a species-specific immunoglobulin-like fold. The structures reveal bona fide nucleic acid binding sites, and nucleic acid binding activities can be detected for NusG from three organisms and for the KOW element alone. A conserved KOW domain is defined as a new class of nucleic acid binding folds. This module is a close structural homolog of tudor protein-protein interaction motifs. Putative protein binding sites for the RNP and KOW domains can be deduced, which differ from the areas implicated in nucleic acid interactions. The results strongly argue that both protein and nucleic acid contacts are important for NusG's functions and that the factor can act as an adaptor mediating indirect protein-nucleic acid associations.
- Max-Planck Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany.
Organizational Affiliation: