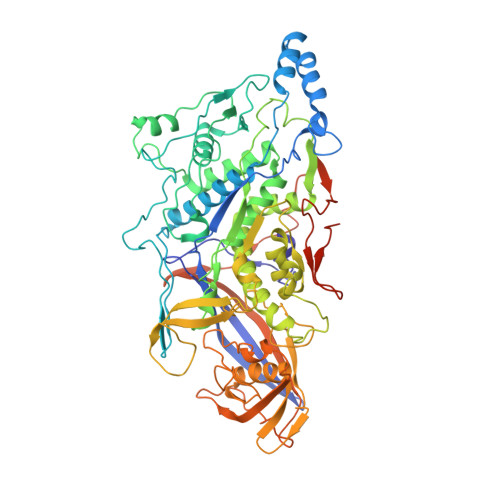
L-A virus at 3.4 A resolution reveals particle architecture and mRNA decapping mechanism.
Naitow, H., Tang, J., Canady, M., Wickner, R.B., Johnson, J.E.(2002) Nat Struct Biol 9: 725-728
- PubMed: 12244300 Search on PubMed
- DOI: https://doi.org/10.1038/nsb844
- Primary Citation Related Structures:
1M1C - PubMed Abstract:
The structure of the yeast L-A virus was determined by X-ray crystallography at 3.4 A resolution. The L-A dsRNA virus is 400 A in diameter and contains a single protein shell of 60 asymmetric dimers of the coat protein, a feature common among the inner protein shells of dsRNA viruses and probably related to their unique mode of transcription and replication. The two identical subunits in each dimer are in non-equivalent environments and show substantially different conformations in specific surface regions. The L-A virus decaps cellular mRNA to efficiently translate its own uncapped mRNA. Our structure reveals a trench at the active site of the decapping reaction and suggests a role for nearby residues in the reaction.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: