Structural Studies of Bacteriophage alpha3 Assembly
Bernal, R.A., Hafenstein, S., Olson, N.H., Bowman, V.D., Chipman, P.R., Baker, T.S., Fane, B.A., Rossmann, M.G.(2003) J Mol Biology 325: 11-24
- PubMed: 12473449 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/s0022-2836(02)01201-9
- Primary Citation Related Structures:
1M06, 1M0F - PubMed Abstract:
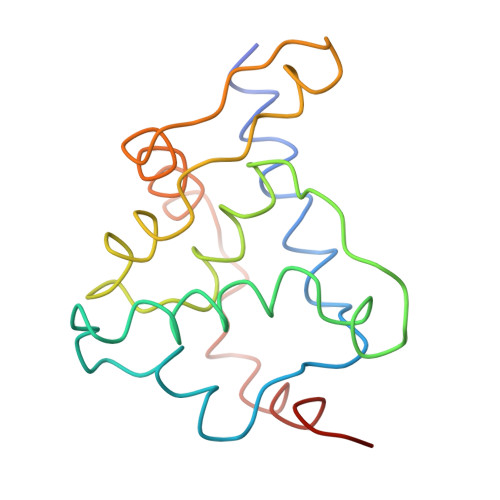
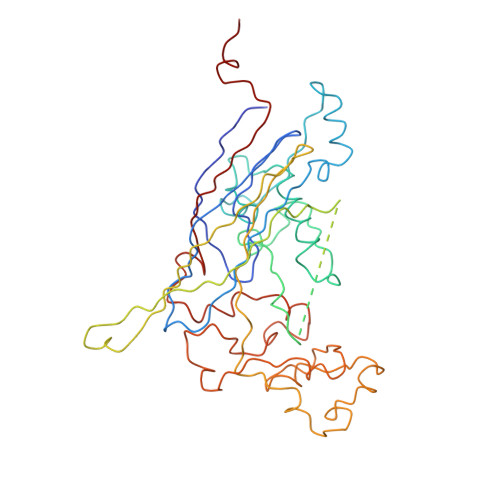
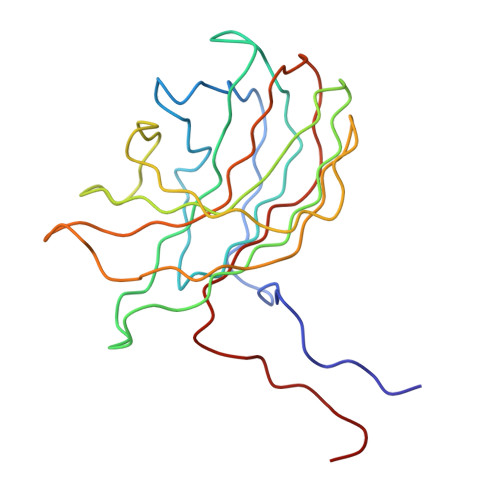
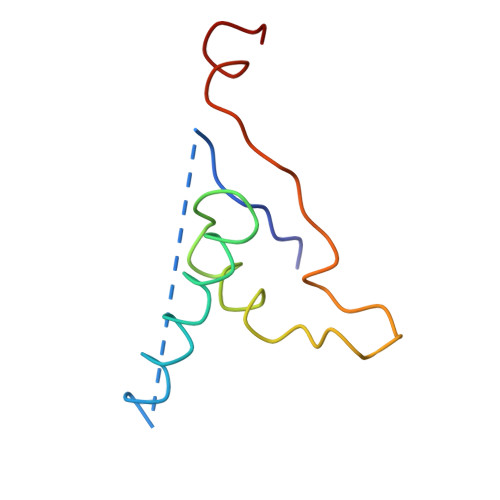
Bacteriophage alpha3 is a member of the Microviridae, a family of small, single-stranded, icosahedral phages that include phiX174. These viruses have an ssDNA genome associated with approximately 12 copies of an H pilot protein and 60 copies of a small J DNA-binding protein. The surrounding capsid consists of 60 F coat proteins decorated with 12 pentameric spikes of G protein. Assembly proceeds via a 108S empty procapsid that requires the external D and internal B scaffolding proteins for its formation. The alpha3 "open" procapsid structural intermediate was determined to 15A resolution by cryo-electron microscopy (cryo-EM). Unlike the phiX174 "closed" procapsid and the infectious virion, the alpha3 open procapsid has 30A wide pores at the 3-fold vertices and 20A wide gaps between F pentamers as a result of the disordering of two helices in the F capsid protein. The large pores are probably used for DNA entry and internal scaffolding protein exit during DNA packaging. Portions of the B scaffolding protein are located at the 5-fold axes under the spike and in the hydrophobic pocket on the inner surface of the capsid. Protein B appears to have autoproteolytic activity that cleaves at an Arg-Phe motif and probably facilitates the removal of the protein through the 30A wide pores. The structure of the alpha3 mature virion was solved to 3.5A resolution by X-ray crystallography and was used to interpret the open procapsid cryo-EM structure. The main differences between the alpha3 and phiX174 virion structures are in the spike and the DNA-binding proteins. The alpha3 pentameric spikes have a rotation of 3.5 degrees compared to those of phiX174. The alpha3 DNA-binding protein, which is shorter by 13 amino acid residues at its amino end when compared to the phiX174 J protein, retains its carboxy-terminal-binding site on the internal surface of the capsid protein. The icosahedrally ordered structural component of the ssDNA appears to be substantially increased in alpha3 compared to phiX174, allowing the building of about 10% of the ribose-phosphate backbone.
- Department of Biological Sciences, Purdue University, 1392 Lilly Hall, West Lafayette, IN 47907-1392, USA.
Organizational Affiliation: