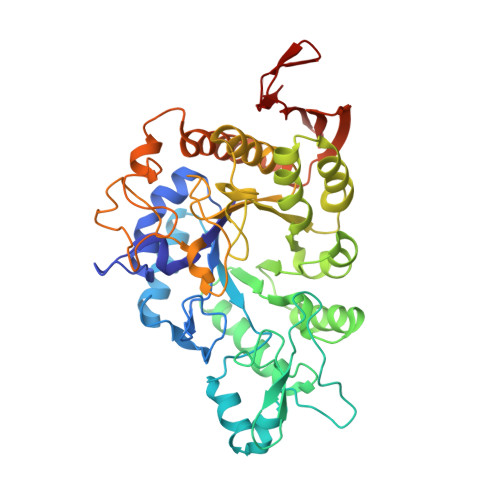
CRYSTAL STRUCTURE OF THERMOTOGA MARITIMA 4-ALPHA-GLUCANOTRANSFERASE AND ITS ACARBOSE COMPLEX: IMPLICATIONS FOR SUBSTRATE SPECIFICITY AND CATALYSIS
Roujeinikova, A., Raasch, C., Sedelnikova, S., Liebl, W., Rice, D.W.(2002) J Mol Biology 321: 149-162
- PubMed: 12139940 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00570-3
- Primary Citation Related Structures:
1LWH, 1LWJ - PubMed Abstract:
4-alpha-Glucanotransferase (GTase) is an essential enzyme in alpha-1,4-glucan metabolism in bacteria and plants. It catalyses the transfer of maltooligosaccharides from an 1,4-alpha-D-glucan molecule to the 4-hydroxyl group of an acceptor sugar molecule. The crystal structures of Thermotoga maritima GTase and its complex with the inhibitor acarbose have been determined at 2.6A and 2.5A resolution, respectively. The GTase structure consists of three domains, an N-terminal domain with the (beta/alpha)(8) barrel topology (domain A), a 65 residue domain, domain B, inserted between strand beta3 and helix alpha6 of the barrel, and a C-terminal domain, domain C, which forms an antiparallel beta-structure. Analysis of the complex of GTase with acarbose has revealed the locations of five sugar-binding subsites (-2 to +3) in the active-site cleft lying between domain B and the C-terminal end of the (beta/alpha)(8) barrel. The structure of GTase closely resembles the family 13 glycoside hydrolases and conservation of key catalytic residues previously identified for this family is consistent with a double-displacement catalytic mechanism for this enzyme. A distinguishing feature of GTase is a pair of tryptophan residues, W131 and W218, which, upon the carbohydrate inhibitor binding, form a remarkable aromatic "clamp" that captures the sugar rings at the acceptor-binding sites +1 and +2. Analysis of the structure of the complex shows that sugar residues occupying subsites from -2 to +2 engage in extensive interactions with the protein, whereas the +3 glucosyl residue makes relatively few contacts with the enzyme. Thus, the structure suggests that four subsites, from -2 to +2, play the dominant role in enzyme-substrate recognition, consistent with the observation that the smallest donor for T.maritima GTase is maltotetraose, the smallest chain transferred is a maltosyl unit and that the smallest residual fragment after transfer is maltose. A close similarity between the structures of GTase and oligo-1,6-glucosidase has allowed the structural features that determine differences in substrate specificity of these two enzymes to be analysed.
- Department of Molecular Biology and Biotechnology, Krebs Institute for Biomolecular Research, The University of Sheffield, England, UK.
Organizational Affiliation: