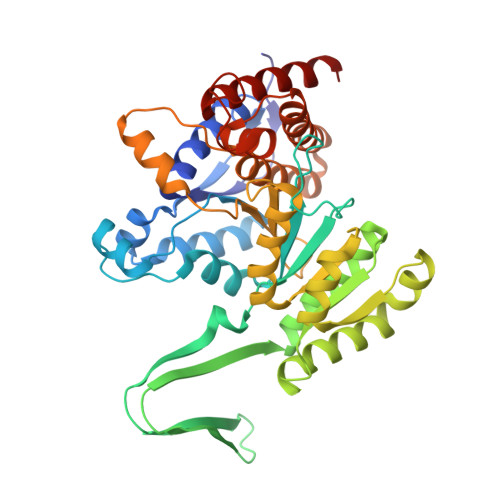
Crystal Structure of Porcine Mitochondrial NADP+-Dependent Isocitrate Dehydrogenase Complexed with Mn2+ and Isocitrate
Ceccarelli, C., Grodsky, N.B., Ariyaratne, N., Colman, R.F., Bahnson, B.J.(2002) J Biological Chem 277: 43454-43462
- PubMed: 12207025
- DOI: https://doi.org/10.1074/jbc.M207306200
- Primary Citation Related Structures:
1LWD - PubMed Abstract:
The crystal structure of porcine heart mitochondrial NADP+-dependent isocitrate dehydrogenase (IDH) complexed with Mn2+ and isocitrate was solved to a resolution of 1.85 A. The enzyme was expressed in Escherichia coli, purified as a fusion protein with maltose binding protein, and cleaved with thrombin to yield homogeneous enzyme. The structure was determined by multiwavelength anomalous diffraction phasing using selenium substitution in the form of selenomethionine as the anomalous scatterer. The porcine NADP+-IDH enzyme is structurally compared with the previously solved structures of IDH from E. coli and Bacillus subtilis that share 16 and 17% identity, respectively, with the mammalian enzyme. The porcine enzyme has a protein fold similar to the bacterial IDH structures with each monomer folding into two domains. However, considerable differences exist between the bacterial and mammalian forms of IDH in regions connecting core secondary structure. Based on the alignment of sequence and structure among the porcine, E. coli, and B. subtilis IDH, a putative phosphorylation site has been identified for the mammalian enzyme. The active site, including the bound Mn2+-isocitrate complex, is highly ordered and, therefore, mechanistically informative. The consensus IDH mechanism predicts that the Mn2+-bound hydroxyl of isocitrate is deprotonated prior to its NADP+-dependent oxidation. The present crystal structure has an active site water that is well positioned to accept the proton and ultimately transfer the proton to solvent through an additional bound water.
- Department of Chemistry and Biochemistry, University of Delaware, Newark, Delaware 19716, USA.
Organizational Affiliation: