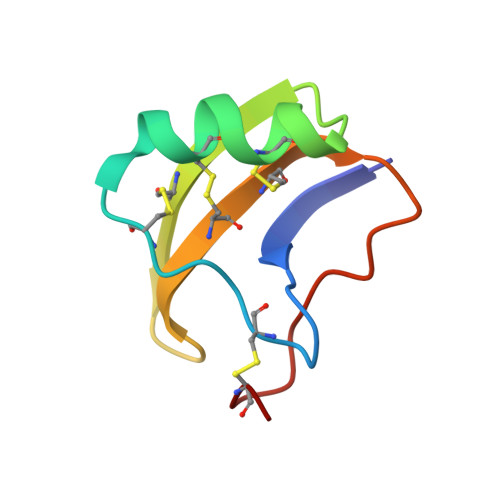
1H-NMR-derived secondary structure and the overall fold of the potent anti-mammal and anti-insect toxin III from the scorpion Leiurus quinquestriatus quinquestriatus.
Landon, C., Cornet, B., Bonmatin, J.M., Kopeyan, C., Rochat, H., Vovelle, F., Ptak, M.(1996) Eur J Biochem 236: 395-404
- PubMed: 8612608 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.1996.00395.x
- Primary Citation Related Structures:
1LQQ - PubMed Abstract:
We describe the secondary structure and the overall fold of toxin III from the venom of the scorpion Leiurus quinquestriatus quinquestriatus determined using two-dimensional-1H-NMR spectroscopy. This protein, which contains 64 amino acids and 4 disulfide bridges, belongs to the long-chain toxin category and is highly toxic to both mammals and insects. The overall fold was determined on the basis of 1208 inter-proton-distance restraints derived from NOE measurements and 90 psi, phi dihedral-angle restraints derived from NOE connectivities and 3JNH-alphaH coupling constants using the HABAS program. This fold, which mainly consists of an alpha-helix packed against a small antiparallel three-stranded beta-sheet, and of several turns and loops, is similar to that of other long-chain scorpion toxins. Aromatic and non-polar residues form several patches on the surface of the protein which alternate with patches of charged and polar residues. Such a topology should be important in the interactions of toxin III with sodium channels in membranes. Two weakly constrained loops introduce some flexibility to the structure which could be related to the activity of this toxin. The central core of toxin III is compared with the cysteine-stabilized alpha beta motif (an alpha-helix connected to a beta-sheet through two disulfide bridges) found in insect defensins and plant thionins. Defensins and thionins are small proteins (approximately 40--50 amino acid residues) containing three or four disulfide bridges, respectively. This comparison confirms that the cysteine-stabilized alpha beta motif is a common core to a number of small proteins from different origins and having different activities.
- Centre de Biophysique Móleculaire (CNRS), Orléans, France.
Organizational Affiliation: