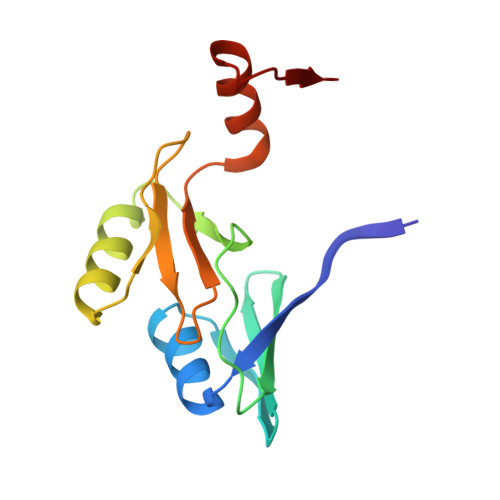
Crystal structure of a genomically encoded fosfomycin resistance protein (FosA) at 1.19 A resolution by MAD phasing off the L-III edge of Tl(+)
Rife, C.L., Pharris, R.E., Newcomer, M.E., Armstrong, R.N.(2002) J Am Chem Soc 124: 11001-11003
- PubMed: 12224946 Search on PubMed
- DOI: https://doi.org/10.1021/ja026879v
- Primary Citation Related Structures:
1LQK, 1LQO, 1LQP - PubMed Abstract:
The fosfomycin resistance protein (FosA) catalyzes the Mn(II)- and K+-dependent addition of glutathione to the oxirane of the antibiotic fosfomycin. The crystal structure of FosA from Pseudomonas aeruginosa was solved at a resolution of 1.19 A by multiwavelength anomalous diffraction at the L-III edge of a Tl+ derivative. The structure solution took advantage of the ability of Tl+ to substitute for K+. The existence of multiple Tl sites in the asymmetric unit suggests that this may be a generally useful technique for phasing protein crystal structures. A 1.35 A resolution structure with phosphate bound in the active site shows that the Mn(II) center has a rare four-coordinate geometry. The structure of the fosfomycin complex at 1.19 A resolution indicates that the Mn(II) center is close to five-coordinate with trigonal bipyramidal geometry and a ligand set consisting of two histidines (H7 and H64) and one phosphonate oxygen occupying the equatorial sites and the carboxylate of E110 at one of the apical sites. The oxirane oxygen of the substrate is located at the other apical site but is 0.2 A beyond the average Mn-O distance for five-coordinate Mn(II). The Mn(II) center is proposed to stabilize the alkoxide in the transition state, while the nearby hydroxyl group of T9 acts as a proton donor in the reaction. The K+ ion located 6.5 A from the Mn(II) appears to help orient the substrate for nucleophilic attack.
- Departments of Biochemistry and Chemistry, Vanderbilt University, Nashville, Tennessee 37232-0146, USA.
Organizational Affiliation: