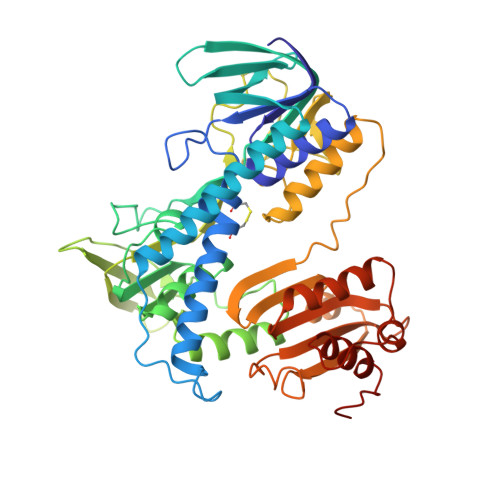
Three-dimensional structure of lipoamide dehydrogenase from Pseudomonas fluorescens at 2.8 A resolution. Analysis of redox and thermostability properties.
Mattevi, A., Obmolova, G., Kalk, K.H., van Berkel, W.J., Hol, W.G.(1993) J Mol Biology 230: 1200-1215
- PubMed: 8487301 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1236
- Primary Citation Related Structures:
1LPF - PubMed Abstract:
The structure of Pseudomonas fluorescens lipoamide dehydrogenase, a dimeric flavoenzyme with a molecular mass of 106,000 daltons, was solved by the molecular replacement method and refined to an R-factor of 19.4% at 2.8 A resolution. The root-mean-square difference from ideal values for bonds and angles is 0.019 A and 3.8 degrees, respectively. The structure is closely related to that of the same flavoprotein from Azotobacter vinelandii. The root-mean-square difference for 932 C alpha atoms is 0.64 A, with 84% sequence identity. The residues in the active site are identical, while 89% of the interface residues are the same in the two enzymes. A few structural variations provide the basis for the differences in thermostability and redox properties between the two homologous proteins. Particularly, in the A. vinelandii molecule a threonine to alanine (T452A) mutation leaves a buried carbonyl oxygen, located at the subunit interface and in proximity of the flavin ring, unpaired to any H-bond donor, probably providing an explanation for the lower stability of the A. vinelandii enzyme with respect to the P. fluorescens enzyme. Six surface loops, which previously could not be accurately positioned in the A. vinelandii structure, are well defined in P. fluorescens lipoamide dehydrogenase. On the basis of the P. fluorescens structure, the six loops could be correctly defined also in the A. vinelandii enzyme. This is an unusual case where similar refinement methodologies applied to two crystal forms of closely related proteins led to electron density maps of substantially different quality. The correct definition of these surface residues is likely to be an essential step for revealing the structural basis of the interactions between lipoamide dehydrogenase and the other members of the pyruvate dehydrogenase multienzyme complex.
- BIOSON Research Institute, Department of Chemistry, University of Groningen, The Netherlands.
Organizational Affiliation: