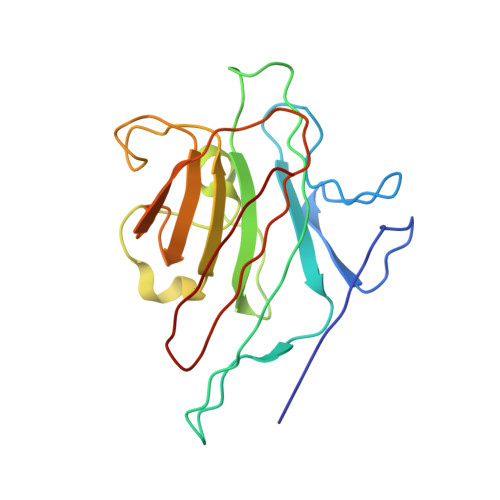
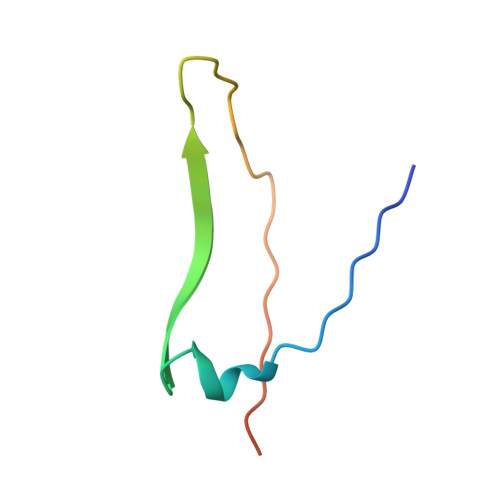
X-ray crystal structure determination and refinement at 1.9 A resolution of isolectin I from the seeds of Lathyrus ochrus.
Bourne, Y., Abergel, C., Cambillau, C., Frey, M., Rouge, P., Fontecilla-Camps, J.C.(1990) J Mol Biology 214: 571-584
- PubMed: 2380988 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(90)90199-V
- Primary Citation Related Structures:
1LOE - PubMed Abstract:
Orthorhombic crystals of isolectin I (LOLI) from the seeds of Lathyrus ochrus were first obtained during the STS 29 space shuttle mission. Subsequently, isostructural crystals were also obtained in the laboratory. They belong to the space group P2(1)2(1)2, with cell dimensions a = 135.84 A, b = 63.12 A and c = 54.54 A with one functional entity, a dimer, in the asymmetric unit (Vm = 2.2 A3/Da). The three-dimensional structure of LOLI, which was solved by the molecular replacement method using a 3 A resolution model of pea lectin, has subsequently been refined by using crystallographic data between 8.0 A and 1.9 A resolution, coupled to molecular dynamics and energy minimization techniques. The conventional R-factor is 0.185 for Fo greater than 1 sigma(Fo). The final model includes 220 well-defined water molecules and has root-mean-square deviations from ideal bond distances and angles of 0.004 A and 3 degrees, respectively. Only slight conformation differences have been found between the two alpha beta monomers. The molecular structure of LOLI, the first to be determined from the genus Lathyrus, is very similar to those of concanavalin A, pea lectin and favin. Differences are confined to the loop regions and beta-chain termini. Comparison of equivalent C alpha atom positions between our final model and the pea lectin structure shows slight differences in the association of the two monomers, which are probably due to the different environments in the crystals. The root-mean-square deviation between C alpha atoms of LOLI and pea lectin is 0.40 A. The metal binding sites are very similar in pea lectin, concanavalin A and LOLI. The sugar-binding sites of LOLI are occupied by four well-ordered water molecules each. The cleavage site for one of the monomers is specially well defined in the final electron density map: the amino group of Glul (alpha) seems to form a salt bridge with the carboxylate group of the terminal Asn181 (beta). A detailed analysis of the difference in crystal packing contacts between pea lectin and LOLI shows that, as might be expected, several of the intermolecular interactions are mediated by residues that correspond to substitutions in the LOLI amino acid sequence.
- Laboratoire de Cristallographie et de Cristallisation des Macromolécules Biologiques, URA 232-CNRS, Faculté de Médecine Secteur-Nord, Marseille, France.
Organizational Affiliation: