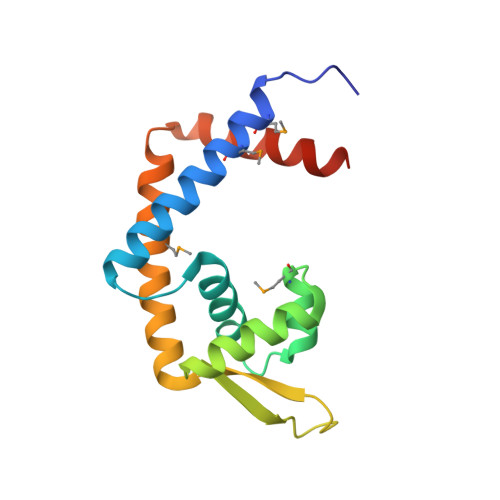
Crystal structure of the MexR repressor of the mexRAB-oprM multidrug efflux operon of Pseudomonas aeruginosa.
Lim, D., Poole, K., Strynadka, N.C.(2002) J Biological Chem 277: 29253-29259
- PubMed: 12034710 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M111381200
- Primary Citation Related Structures:
1LNW - PubMed Abstract:
MexR is a member of the MarR family of bacterial transcriptional regulators and is the repressor for the MexAB-OprM operon, which encodes a tripartite multidrug efflux system in Pseudomonas aeruginosa. Mutations in MexR result in increased resistance to multiple antibiotics due to overexpression of this efflux system. We have determined the crystal structure of MexR to 2.1-A resolution in the absence of effector. The four copies of the MexR dimer in the asymmetric unit are observed in multiple conformations. Analysis of these conformational states in the context of a model of the MexR-DNA complex proposed in this study suggests that an effector-induced conformational change may inhibit DNA binding by reducing the spacing of the DNA binding domains. The inhibited conformation is exhibited by one of the four MexR dimers, which contains an ordered C-terminal tail from a neighboring monomer inserted between its DNA binding domains and which we propose may resemble the MexR-effector complex. Our results indicate that MexR may differ from the other described member of this family, MarR, in the nature of its effector, mode of DNA binding, and mechanism of regulation.
- Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada.
Organizational Affiliation: