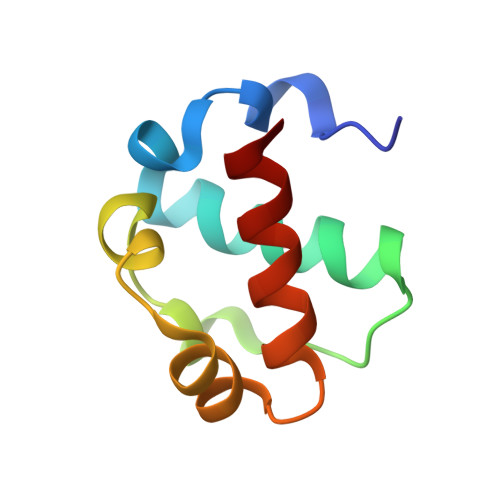
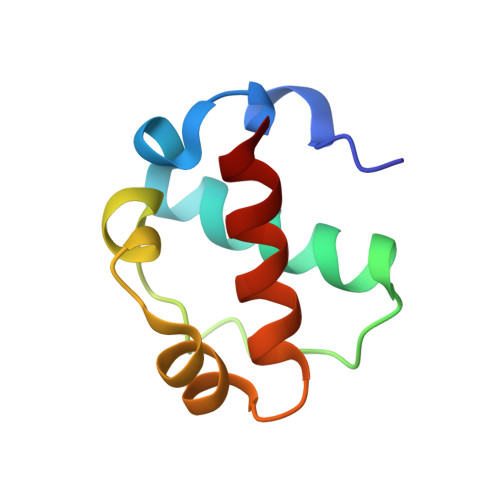
Native interface of the SAM domain polymer of TEL.
Tran, H.H., Kim, C.A.(2002) BMC Struct Biol 2: 5-5
- PubMed: 12193272 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-2-5
- Primary Citation Related Structures:
1LKY - PubMed Abstract:
TEL is a transcriptional repressor containing a SAM domain that forms a helical polymer. In a number of hematologic malignancies, chromosomal translocations lead to aberrant fusions of TEL-SAM to a variety of other proteins, including many tyrosine kinases. TEL-SAM polymerization results in constitutive activation of the tyrosine kinase domains to which it becomes fused, leading to cell transformation. Thus, inhibitors of TEL-SAM self-association could abrogate transformation in these cells. In previous work, we determined the structure of a mutant TEL-SAM polymer bearing a Val to Glu substitution in center of the subunit interface. It remained unclear how much the mutation affected the architecture of the polymer, however. Here we determine the structure of the native polymer interface. To accomplish this goal, we introduced mutations that block polymer extension, producing a heterodimer with a wild-type interface. We find that the structure of the wild-type polymer interface is quite similar to the mutant structure determined previously. With the structure of the native interface, it is possible to evaluate the potential for developing therapeutic inhibitors of the interaction. We find that the interacting surfaces of the protein are relatively flat, containing no obvious pockets for the design of small molecule inhibitors. Our results confirm the architecture of the TEL-SAM polymer proposed previously based on a mutant structure. The fact that the interface contains no obvious potential binding pockets suggests that it may be difficult to find small molecule inhibitors to treat malignancies in this way.
- Department of Chemistry and Biochemistry, UCLA Laboratory of Structural Biology and Molecular Medicine, Molecular Biology Institute, 90095-1570, USA. htran@mbi.ucla.edu
Organizational Affiliation: