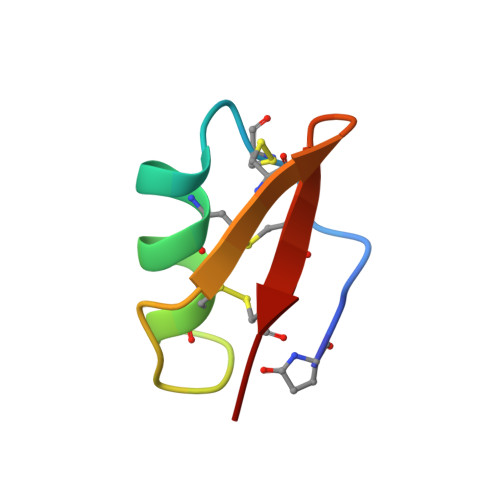
Solution structure of potassium channel-inhibiting scorpion toxin Lq2.
Renisio, J.G., Lu, Z., Blanc, E., Jin, W., Lewis, J.H., Bornet, O., Darbon, H.(1999) Proteins 34: 417-426
- PubMed: 10081954
- DOI: https://doi.org/10.1002/(sici)1097-0134(19990301)34:4<417::aid-prot1>3.0.co;2-r
- Primary Citation of Related Structures:
1LIR - PubMed Abstract:
Lq2 is a unique scorpion toxin. Acting from the extracellular side, Lq2 blocks the ion conduction pore in not only the voltage- and Ca2+ -activated channels, but also the inward-rectifier K+ channels. This finding argues that the three-dimensional structures of the pores in these K+ channels are similar. However, the amino acid sequences that form the external part of the pore are minimally conserved among the various classes of K+ channels. Because Lq2 can bind to all the three classes of K+ channels, we can use Lq2 as a structural probe to examine how the non-conserved pore-forming sequences are arranged in space to form similar pore structures. In the present study, we determined the three-dimensional structure of Lq2 using nuclear magnetic resonance (NMR) techniques. Lq2 consists of an alpha-helix (residues S10 to L20) and a beta-sheet, connected by an alphabeta3 loop (residues N22 to N24). The beta-sheet has two well-defined anti-parallel strands (residues G26 to M29 and residues K32 to C35), which are connected by a type I' beta-turn centered between residues N30 and K31. The N-terminal segment (residues Z1 to T8) appears to form a quasi-third strand of the beta-sheet.
- Architecture et Fonction des Macromolécules Biologiques, Centre National de la Recherche Scientifique, UPR 9039, Marseille, France.
Organizational Affiliation: