A New Zinc Binding Fold Underlines the Versatility of Zinc Binding Modules in Protein Evolution
Sharpe, B.K., Matthews, J.M., Kwan, A.H.Y., Newton, A., Gell, D.A., Crossley, M., Mackay, J.P.(2002) Structure 10: 639-648
- PubMed: 12015147
- DOI: https://doi.org/10.1016/s0969-2126(02)00757-8
- Primary Citation Related Structures:
1LIQ - PubMed Abstract:
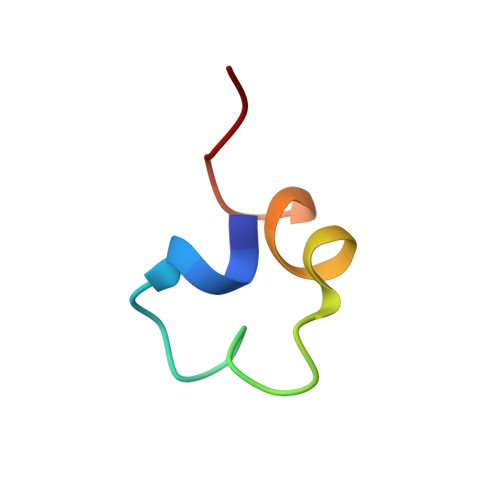
Many different zinc binding modules have been identified. Their abundance and variety suggests that the formation of zinc binding folds might be relatively common. We have determined the structure of CH1(1), a 27-residue peptide derived from the first cysteine/histidine-rich region (CH1) of CREB binding protein (CBP). This peptide forms a highly ordered zinc-dependent fold that is distinct from known folds. The structure differs from a subsequently determined structure of a larger region from the CH3 region of CBP, and the CH1(1) fold probably represents a nonphysiologically active form. Despite this, the fold is thermostable and tolerant to both multiple alanine mutations and changes in the zinc-ligand spacing. Our data support the idea that zinc binding domains may arise frequently. Additionally, such structures may prove useful as scaffolds for protein design, given their stability and robustness.
- Department of Biochemistry, University of Sydney, New South Wales 2006, Australia.
Organizational Affiliation: