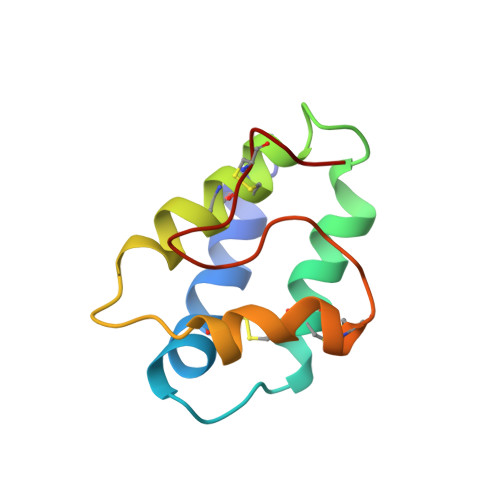
Structure in solution of a four-helix lipid binding protein.
Heinemann, B., Andersen, K.V., Nielsen, P.R., Bech, L.M., Poulsen, F.M.(1996) Protein Sci 5: 13-23
- PubMed: 8771192 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560050103
- Primary Citation Related Structures:
1LIP - PubMed Abstract:
Because of the low solubility of lipids in water, intercellular and intracellular pathways of lipid transfer are necessary, e.g., for membrane formation. The mechanism by which lipids in vivo are transported from their site of biogenesis (endoplasmatic reticulum and the chloroplasts) to their place of action is unknown. Several small plant proteins with the ability to mediate transfer of radiolabeled phospholipids in vitro from liposomal donor membranes to mitochondrial and chloroplast acceptor membranes have been isolated, and a protein with this ability, the nonspecific lipid transfer protein (nsLTP) isolated from barley seeds (bLTP), has been studied here. The structure and the protein lipid interactions of lipid transfer proteins are relevant for the understanding of their function, and here we present the three-dimensional structure in solution of bLTP as determined by NMR spectroscopy. The 1H NMR spectrum of the 91-residue protein was assigned for more than 97% of the protein 1H atoms, and the structure was calculated on the basis of 813 distance restraints from 1H-1H nuclear Overhauser effects, four disulfide bond restraints, from dihedral angle restraints for 66 phi-angles, 61 chi 1 angles, and 2 chi 2 angles, and from 31 sets of hydrogen bond restraints. The solution structure of bLTP consists of four well-defined alpha-helices A-D (A, Cys 3-Gly 19; B, Gly 25-Ala 38; C, Arg 44-Gly 57; D, Leu 63-Cys 73), separated by three short loops that are less well defined and concluded by a well defined C-terminal peptide segment with no observable regular secondary structure. For the 17 structures that are used to represent the solution structure of bLTP, the RMS deviation to an average structure is 0.63 A +/- 0.04 A for backbone atoms and 0.93 A +/- 0.06 A for all heavy atoms. The secondary structure elements and their locations in the sequence resemble those of nsLTP from two other plant species, wheat and maize, whose structures were previously determined (Gincel E et al, 1995, Eur J Biochem 226:413-422; Shin DH et al, 1995, Structure 3:189-199). In bLTP, the residues analogous to those in maize nsLTP that constitute the palmitate binding site are forming a similar hydrophobic cavity and a potential acyl group binding site. Analysis of the solution structure of bLTP and bLTP in complex with a ligand might provide information on the conformational changes in the protein upon ligand binding and subsequently provide information on the mode of ligand uptake and release. In this work, we hope to establish a foundation for further work of determining the solution structure of bLTP in complex with palmitoyl coenzyme A, which is a suitable ligand, and subsequently to outline the mode of ligand binding.
- Carlsberg Laboratorium, Kemisk Afdeling, Copenhagen, Denmark.
Organizational Affiliation: