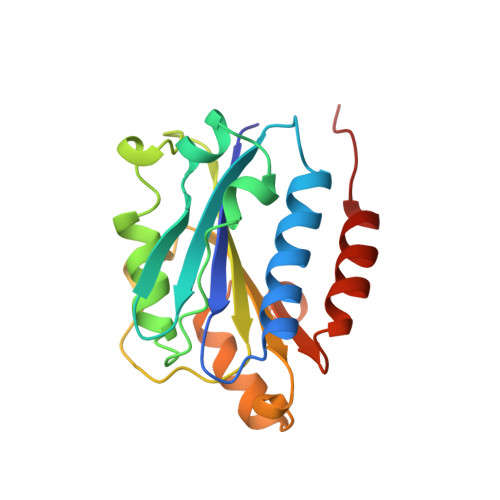
Crystal structure of the I-domain from the CD11a/CD18 (LFA-1, alpha L beta 2) integrin.
Qu, A., Leahy, D.J.(1995) Proc Natl Acad Sci U S A 92: 10277-10281
- PubMed: 7479767 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.22.10277
- Primary Citation Related Structures:
1LFA - PubMed Abstract:
We report the 1.8-A crystal structure of the CD11a I-domain with bound manganese ion. The CD11a I-domain contains binding sites for intercellular adhesion molecules 1 and 3 and can exist in both low- and high-affinity states. The metal-bound form reported here is likely to represent a high-affinity state. The CD11a I-domain structure reveals a strained hydrophobic ridge adjacent to the bound metal ion that may serve as a ligand-binding surface and is likely to rearrange in the absence of bound metal ion. The CD11a I-domain is homologous to domains found in von Willebrand factor, and mapping of mutations found in types 2a and 2b von Willebrand disease onto this structure allows consideration of the molecular basis of these forms of the disease.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: