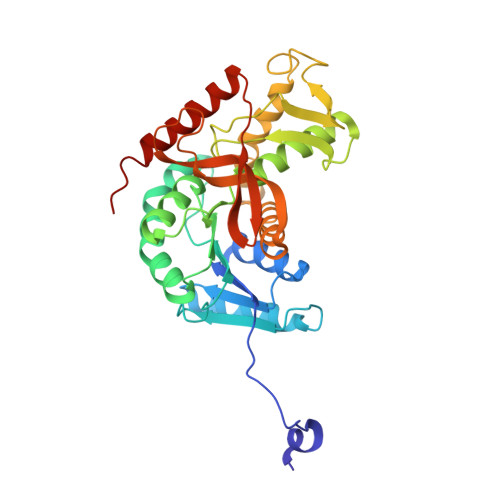
Refined crystal structure of dogfish M4 apo-lactate dehydrogenase.
Abad-Zapatero, C., Griffith, J.P., Sussman, J.L., Rossmann, M.G.(1987) J Mol Biology 198: 445-467
- PubMed: 3430615 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(87)90293-2
- Primary Citation Related Structures:
1LDM, 6LDH, 8LDH - PubMed Abstract:
The crystal structure of M4 apo-lactate dehydrogenase from the spiny dogfish (Squalus acanthius) was initially refined by a constrained-restrained, and subsequently restrained, least-squares technique. The final structure contained 286 water molecules and two sulfate ions per subunit and gave an R-factor of 0.202 for difraction data between 8.0 and 2.0 A resolution. The upper limit for the co-ordinate accuracy of the atoms was estimated to be 0.25 A. The elements of secondary structure of the refined protein have not changed from those described previously, except for the appearance of a one-and-a-half turn 3(10) helix immediately after beta J. There is also a short segment of 3(10) helix between beta C and beta D in the part of the chain that connects the two beta alpha beta alpha beta units of the six-stranded parallel sheet (residues Tyr83 to Ala87). Examination of the interactions among the different elements of secondary structure by means of a surface accessibility algorithm supports the four structural clusters in the subunit. The first of the two sulfate ions is in the active site and occupies a cavity near the essential His195. Its nearest protein ligands are Arg171, Asp168 and Asn140. The second sulfate ion is located near the P-axis subunit interface. It is liganded by His188 and Arg173. These two residues are conserved in bacterial lactate dehydrogenase and form part of the fructose 1,6-bisphosphate effector binding site. Two other data sets in which one (collected at pH 7.8) or both (collected at pH 6.0) sulfate ions were replaced by citrate ions were also analyzed. Five cycles of refinement with respect to the pH 6.0 data (25 to 2.8 A resolution) resulted in an R value of 0.191. Only water molecules occupy the subunit boundary anion binding site at pH 7.8. The amino acid sequence was found to be in poor agreement with (2Fobs-Fcalc) electron density maps for the peptide between residues 207 and 211. The original sequence WNALKE was replaced by NVASIK. The essential His195 is hydrogen bonded to Asp168 on one side and Asn140 on the other. The latter residue is part of a turn that contains the only cis peptide bond of the structure at Pro141. The "flexible loop" (residues 97 to 123), which folds down over the active center in ternary complexes of the enzyme with substrate and coenzyme, has a well-defined structure. Analysis of the environment of Tyr237 suggests how its chemical modification inhibits the enzyme.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907.
Organizational Affiliation: